Abstract
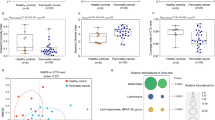
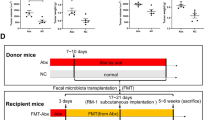
Pancreatic cancer is a lethal cancer with aggressive and invasive characteristics. By the time it is diagnosed, patients already have tumors extended to other organs and show extremely low survival rates. The gut microbiome is known to be associated with many diseases and its imbalance affects the pathogenesis of pancreatic cancer. In this study, we established an orthotopic, patient-derived xenograft model to identify how the gut microbiome is linked to pancreatic ductal adenocarcinoma (PDAC). Using the 16S rDNA metagenomic sequencing, we revealed that the levels of Alistipes onderdonkii and Roseburia hominis decreased in the gut microbiome of the PDAC model. To explore the crosstalk between the two bacteria and PDAC cells, we collected the supernatant of the bacteria or cancer cell culture medium and treated it in a cross manner. While the cancer cell medium did not affect bacterial growth, we observed that the A. onderdonkii medium suppressed the growth of the pancreatic primary cancer cells. Using the bromodeoxyuridine/7-amino-actinomycin D (BrdU/7-AAD) staining assay, we confirmed that the A. onderdonkii medium inhibited the proliferation of the pancreatic primary cancer cells. Furthermore, RNA-seq analysis revealed that the A. onderdonkii medium induced unique transcriptomic alterations in the PDAC cells, compared to the normal pancreatic cells. Altogether, our data suggest that the reduction in the A. onderdonkii in the gut microbiome provides a proliferation advantage to the pancreatic cancer cells.
Graphical abstract

Key points
• Metagenome analysis of pancreatic cancer model reveals A. onderdonkii downregulation.
• A. onderdonkii culture supernatant suppressed the proliferation of pancreatic cancer cells.
• RNA seq data reveals typical gene expression changes induced by A. onderdonkii.






Similar content being viewed by others
Data availability
For metagenome data, SRA records will be accessible with the following link https://www.ncbi.nlm.nih.gov/sra/PRJNA733811.
For other raw data, Supplementary material is available online.
References
Amieva M, Peek RM Jr (2016) Pathobiology of Helicobacter pylori-induced gastric cancer. Gastroenterology 150(1):64–78. https://doi.org/10.1053/j.gastro.2015.09.004
Anders S, Pyl PT, Huber W (2015) HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics 31(2):166–169. https://doi.org/10.1093/bioinformatics/btu638
Anselme K, Davidson P, Popa AM, Giazzon M, Liley M, Ploux L (2010) The interaction of cells and bacteria with surfaces structured at the nanometre scale. Acta Biomater 6(10):3824–3846. https://doi.org/10.1016/j.actbio.2010.04.001
Arumugam M, Agarwal A, Arya MC, Ahmed Z (2013) Influence of nitrogen sources on biomass productivity of microalgae Scenedesmus bijugatus. Bioresour Technol 131:246–249. https://doi.org/10.1016/j.biortech.2012.12.159
Biedermann L, Zeitz J, Mwinyi J, Sutter-Minder E, Rehman A, Ott SJ, Steurer-Stey C, Frei A, Frei P, Scharl M, Loessner MJ, Vavricka SR, Fried M, Schreiber S, Schuppler M, Rogler G (2013) Smoking cessation induces profound changes in the composition of the intestinal microbiota in humans. PLoS ONE 8(3):e59260. https://doi.org/10.1371/journal.pone.0059260
Chan AA, Bashir M, Rivas MN, Duvall K, Sieling PA, Pieber TR, Vaishampayan PA, Love SM, Lee DJ (2016) Characterization of the microbiome of nipple aspirate fluid of breast cancer survivors. Sci Rep 6:28061. https://doi.org/10.1038/srep28061
Chapman PB, Hauschild A, Robert C, Haanen JB, Ascierto P, Larkin J, Dummer R, Garbe C, Testori A, Maio M, Hogg D, Lorigan P, Lebbe C, Jouary T, Schadendorf D, Ribas A, O'Day SJ, Sosman JA, Kirkwood JM, Eggermont AM, Dreno B, Nolop K, Li J, Nelson B, Hou J, Lee RJ, Flaherty KT, McArthur GA, Group B-S (2011) Improved survival with vemurafenib in melanoma with BRAF V600E mutation. N Engl J Med 364(26):2507-16. https://doi.org/10.1056/NEJMoa1103782
Costa E, Teixidó N, Usall J, Atarés E, Viñas I (2002) The effect of nitrogen and carbon sources on growth of the biocontrol agent Pantoea agglomerans strain CPA-2. Lett Appl Microbiol 35(2):117–120. https://doi.org/10.1046/j.1472-765x.2002.01133.x
Dal Molin M, Zhang M, de Wilde RF, Ottenhof NA, Rezaee N, Wolfgang CL, Blackford A, Vogelstein B, Kinzler KW, Papadopoulos N, Hruban RH, Maitra A, Wood LD (2015) Very long-term survival following resection for pancreatic cancer is not explained by commonly mutated genes: results of whole-exome sequencing analysis. Clin Cancer Res 21(8):1944–1950. https://doi.org/10.1158/1078-0432.CCR-14-2600
Dalile B, Van Oudenhove L, Vervliet B, Verbeke K (2019) The role of short-chain fatty acids in microbiota-gut-brain communication. Nat Rev Gastroenterol Hepatol 16(8):461–478. https://doi.org/10.1038/s41575-019-0157-3
Deng M, Zhang W, Yuan L, Tan J, Chen Z (2020) HIF-1a regulates hypoxia-induced autophagy via translocation of ANKRD37 in colon cancer. Exp Cell Res 395(1):112175. https://doi.org/10.1016/j.yexcr.2020.112175
DeRose YS, Wang G, Lin YC, Bernard PS, Buys SS, Ebbert MT, Factor R, Matsen C, Milash BA, Nelson E, Neumayer L, Randall RL, Stijleman IJ, Welm BE, Welm AL (2011) Tumor grafts derived from women with breast cancer authentically reflect tumor pathology, growth, metastasis and disease outcomes. Nat Med 17(11):1514–1520. https://doi.org/10.1038/nm.2454
Desai NV, Sliesoraitis S, Hughes SJ, Trevino JG, Zlotecki RA, Ivey AM, George TJ Jr (2015) Multidisciplinary neoadjuvant management for potentially curable pancreatic cancer. Cancer Med 4(8):1224–1239. https://doi.org/10.1002/cam4.444
Dobin A, Davis CA, Schlesinger F, Drenkow J, Zaleski C, Jha S, Batut P, Chaisson M, Gingeras TR (2013) STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29(1):15–21. https://doi.org/10.1093/bioinformatics/bts635
Farhana L, Banerjee HN, Verma M, Majumdar APN (2018) Role of microbiome in carcinogenesis process and epigenetic regulation of colorectal cancer. Methods Mol Biol 1856:35–55. https://doi.org/10.1007/978-1-4939-8751-1_3
Feng Q, Liang S, Jia H, Stadlmayr A, Tang L, Lan Z, Zhang D, **a H, Xu X, Jie Z, Su L, Li X, Li X, Li J, **ao L, Huber-Schonauer U, Niederseer D, Xu X, Al-Aama JY, Yang H, Wang J, Kristiansen K, Arumugam M, Tilg H, Datz C, Wang J (2015) Gut microbiome development along the colorectal adenoma-carcinoma sequence. Nat Commun 6:6528. https://doi.org/10.1038/ncomms7528
Flemer B, Lynch DB, Brown JM, Jeffery IB, Ryan FJ, Claesson MJ, O’Riordain M, Shanahan F, O’Toole PW (2017) Tumour-associated and non-tumour-associated microbiota in colorectal cancer. Gut 66(4):633–643. https://doi.org/10.1136/gutjnl-2015-309595
Frank DN, St Amand AL, Feldman RA, Boedeker EC, Harpaz N, Pace NR (2007) Molecular-phylogenetic characterization of microbial community imbalances in human inflammatory bowel diseases. Proc Natl Acad Sci U S A 104(34):13780–13785. https://doi.org/10.1073/pnas.0706625104
Fujisaka S, Avila-Pacheco J, Soto M, Kostic A, Dreyfuss JM, Pan H, Ussar S, Altindis E, Li N, Bry L, Clish CB, Kahn CR (2018) Diet, genetics, and the gut microbiome drive dynamic changes in plasma metabolites. Cell Rep 22(11):3072–3086. https://doi.org/10.1016/j.celrep.2018.02.060
Gill SR, Pop M, Deboy RT, Eckburg PB, Turnbaugh PJ, Samuel BS, Gordon JI, Relman DA, Fraser-Liggett CM, Nelson KE (2006) Metagenomic analysis of the human distal gut microbiome. Science 312(5778):1355–1359. https://doi.org/10.1126/science.1124234
Goodman B, Gardner H (2018) The microbiome and cancer. J Pathol 244(5):667–676. https://doi.org/10.1002/path.5047
Goodwin AC, Destefano Shields CE, Wu S, Huso DL, Wu X, Murray-Stewart TR, Hacker-Prietz A, Rabizadeh S, Woster PM, Sears CL, Casero RA Jr (2011) Polyamine catabolism contributes to enterotoxigenic Bacteroides fragilis-induced colon tumorigenesis. Proc Natl Acad Sci U S A 108(37):15354–15359. https://doi.org/10.1073/pnas.1010203108
Gori S, Inno A, Belluomini L, Bocus P, Bisoffi Z, Russo A, Arcaro G (2019) Gut microbiota and cancer: how gut microbiota modulates activity, efficacy and toxicity of antitumoral therapy. Crit Rev Oncol Hematol 143:139–147. https://doi.org/10.1016/j.critrevonc.2019.09.003
Guarner F, Malagelada J-R (2003) Gut flora in health and disease. The Lancet 361(9356):512–519. https://doi.org/10.1016/s0140-6736(03)12489-0
Half E, Keren N, Reshef L, Dorfman T, Lachter I, Kluger Y, Reshef N, Knobler H, Maor Y, Stein A, Konikoff FM, Gophna U (2019) Fecal microbiome signatures of pancreatic cancer patients. Sci Rep 9(1):16801. https://doi.org/10.1038/s41598-019-53041-4
Humblot C, Murkovic M, Rigottier-Gois L, Bensaada M, Bouclet A, Andrieux C, Anba J, Rabot S (2007) Beta-glucuronidase in human intestinal microbiota is necessary for the colonic genotoxicity of the food-borne carcinogen 2-amino-3-methylimidazo[4,5-f]quinoline in rats. Carcinogenesis 28(11):2419–2425. https://doi.org/10.1093/carcin/bgm170
Jain T, Dudeja V (2021) The war against pancreatic cancer in 2020 - advances on all fronts. Nat Rev Gastroenterol Hepatol 18(2):99–100. https://doi.org/10.1038/s41575-020-00410-4
Jung J, Lee CH, Seol HS, Choi YS, Kim E, Lee EJ, Rhee JK, Singh SR, Jun ES, Han B, Hong SM, Kim SC, Chang S (2016) Generation and molecular characterization of pancreatic cancer patient-derived xenografts reveals their heterologous nature. Oncotarget 7(38):62533–62546. https://doi.org/10.18632/oncotarget.11530
Kazmierczak BI, Mostov K, Engel JN (2001) Interaction of bacterial pathogens with polarized epithelium. Annu Rev Microbiol 55:407–435. https://doi.org/10.1146/annurev.micro.55.1.407
Klampfer L (2008) The role of signal transducers and activators of transcription in colon cancer. Front Biosci 13(1):2888–2899. https://doi.org/10.2741/2893
Lavu H, Nowcid LJ, Klinge MJ, Mahendraraj K, Grenda DR, Sauter PK, Rosato EL, Kennedy EP, Yeo CJ (2011) Reoperative completion pancreatectomy for suspected malignant disease of the pancreas. J Surg Res 170(1):89–95. https://doi.org/10.1016/j.jss.2011.04.050
Lee SK, Moon JW, Lee YW, Lee JO, Kim SJ, Kim N, Kim J, Kim HS, Park SH (2015) The effect of high glucose levels on the hypermethylation of protein phosphatase 1 regulatory subunit 3C (PPP1R3C) gene in colorectal cancer. J Genet 94(1):75–85. https://doi.org/10.1007/s12041-015-0492-2
Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, Marth G, Abecasis G, Durbin R (2009) The sequence alignment/map format and SAMtools. Bioinformatics 25(16):2078–2079. https://doi.org/10.1093/bioinformatics/btp352
Li J, Sung CY, Lee N, Ni Y, Pihlajamaki J, Panagiotou G, El-Nezami H (2016) Probiotics modulated gut microbiota suppresses hepatocellular carcinoma growth in mice. Proc Natl Acad Sci U S A 113(9):E1306–E1315. https://doi.org/10.1073/pnas.1518189113
Litman-Zawadzka A, Lukaszewicz-Zajac M, Mroczko B (2019) Novel potential biomarkers for pancreatic cancer - a systematic review. Adv Med Sci 64(2):252–257. https://doi.org/10.1016/j.advms.2019.02.004
Machiels K, Joossens M, Sabino J, De Preter V, Arijs I, Eeckhaut V, Ballet V, Claes K, Van Immerseel F, Verbeke K, Ferrante M, Verhaegen J, Rutgeerts P, Vermeire S (2014) A decrease of the butyrate-producing species Roseburia hominis and Faecalibacterium prausnitzii defines dysbiosis in patients with ulcerative colitis. Gut 63(8):1275–1283. https://doi.org/10.1136/gutjnl-2013-304833
Malaney P, Nicosia SV, Dave V (2014) One mouse, one patient paradigm: new avatars of personalized cancer therapy. Cancer Lett 344(1):1–12. https://doi.org/10.1016/j.canlet.2013.10.010
Manichanh C, Rigottier-Gois L, Bonnaud E, Gloux K, Pelletier E, Frangeul L, Nalin R, Jarrin C, Chardon P, Marteau P, Roca J, Dore J (2006) Reduced diversity of faecal microbiota in Crohn’s disease revealed by a metagenomic approach. Gut 55(2):205–211. https://doi.org/10.1136/gut.2005.073817
Martin M (2011) Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J 17(1):10–12. https://doi.org/10.14806/ej.17.1.200
Meraz IM, Majidi M, Meng F, Shao R, Ha MJ, Neri S, Fang B, Lin SH, Tinkey PT, Shpall EJ, Morris J, Roth JA (2019) An improved patient-derived xenograft humanized mouse model for evaluation of lung cancer immune responses. Cancer Immunol Res 7(8):1267–1279. https://doi.org/10.1158/2326-6066.CIR-18-0874
Moschen AR, Gerner RR, Wang J, Klepsch V, Adolph TE, Reider SJ, Hackl H, Pfister A, Schilling J, Moser PL, Kempster SL, Swidsinski A, Orth Holler D, Weiss G, Baines JF, Kaser A, Tilg H (2016) Lipocalin 2 protects from inflammation and tumorigenesis associated with gut microbiota alterations. Cell Host Microbe 19(4):455–469. https://doi.org/10.1016/j.chom.2016.03.007
Ohnishi S, Futamura M, Kamino H, Nakamura Y, Kitamura N, Miyamoto Y, Miyamoto T, Shinogi D, Goda O, Arakawa H (2010) Identification of NEEP21, encoding neuron-enriched endosomal protein of 21 kDa, as a transcriptional target of tumor suppressor p53. Int J Oncol 37(5):1133–1141. https://doi.org/10.3892/ijo_00000765
O’Rourke F, Kempf VAJ (2019) Interaction of bacteria and stem cells in health and disease. FEMS Microbiol Rev 43(2):162–180. https://doi.org/10.1093/femsre/fuz003
Parker BJ, Wearsch PA, Veloo ACM, Rodriguez-Palacios A (2020) The genus Alistipes: gut bacteria with emerging implications to inflammation, cancer, and mental health. Front Immunol 11:906. https://doi.org/10.3389/fimmu.2020.00906
Patterson AM, Mulder IE, Travis AJ, Lan A, Cerf-Bensussan N, Gaboriau-Routhiau V, Garden K, Logan E, Delday MI, Coutts AGP, Monnais E, Ferraria VC, Inoue R, Grant G, Aminov RI (2017) Human gut symbiont Roseburia hominis promotes and regulates innate immunity. Front Immunol 8:1166. https://doi.org/10.3389/fimmu.2017.01166
Posselt G, Backert S, Wessler S (2013) The functional interplay of Helicobacter pylori factors with gastric epithelial cells induces a multi-step process in pathogenesis. Cell Commun Signal 11:77. https://doi.org/10.1186/1478-811X-11-77
Pushalkar S, Hundeyin M, Daley D, Zambirinis CP, Kurz E, Mishra A, Mohan N, Aykut B, Usyk M, Torres LE, Werba G, Zhang K, Guo Y, Li Q, Akkad N, Lall S, Wadowski B, Gutierrez J, Kochen Rossi JA, Herzog JW, Diskin B, Torres-Hernandez A, Leinwand J, Wang W, Taunk PS, Savadkar S, Janal M, Saxena A, Li X, Cohen D, Sartor RB, Saxena D, Miller G (2018) The pancreatic cancer microbiome promotes oncogenesis by induction of innate and adaptive immune suppression. Cancer Discov 8(4):403–416. https://doi.org/10.1158/2159-8290.CD-17-1134
Quan Y, Song K, Zhang Y, Zhu C, Shen Z, Wu S, Luo W, Tan B, Yang Z, Wang X (2018) Roseburia intestinalis-derived flagellin is a negative regulator of intestinal inflammation. Biochem Biophys Res Commun 501(3):791–799. https://doi.org/10.1016/j.bbrc.2018.05.075
Reyal F, Guyader C, Decraene C, Lucchesi C, Auger N, Assayag F, De Plater L, Gentien D, Poupon MF, Cottu P, De Cremoux P, Gestraud P, Vincent-Salomon A, Fontaine JJ, Roman-Roman S, Delattre O, Decaudin D, Marangoni E (2012) Molecular profiling of patient-derived breast cancer xenografts. Breast Cancer Res 14(1):R11. https://doi.org/10.1186/bcr3095
Ríos-Covián D, Ruas-Madiedo P, Margolles A, Gueimonde M, de Los Reyes-Gavilán CG, Salazar N (2016) Intestinal short chain fatty acids and their link with diet and human health. Front Microbiol 17(7):185. https://doi.org/10.3389/fmicb.2016.00185
Riquelme E, Zhang Y, Zhang L, Montiel M, Zoltan M, Dong W, Quesada P, Sahin I, Chandra V, San Lucas A, Scheet P, Xu H, Hanash SM, Feng L, Burks JK, Do KA, Peterson CB, Nejman D, Tzeng CD, Kim MP, Sears CL, Ajami N, Petrosino J, Wood LD, Maitra A, Straussman R, Katz M, White JR, Jenq R, Wargo J, McAllister F (2019) Tumor microbiome diversity and composition influence pancreatic cancer outcomes. Cell 178(4):795–806 e12. https://doi.org/10.1016/j.cell.2019.07.008
Robinson MD, McCarthy DJ, Smyth GK (2010) edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26(1):139–140. https://doi.org/10.1093/bioinformatics/btp616
Sartor RB, Wu GD (2017) Roles for intestinal bacteria, viruses, and fungi in pathogenesis of inflammatory bowel diseases and therapeutic approaches. Gastroenterology 152(2):327–339 e4. https://doi.org/10.1053/j.gastro.2016.10.012
Sethi V, Kurtom S, Tarique M, Lavania S, Malchiodi Z, Hellmund L, Zhang L, Sharma U, Giri B, Garg B, Ferrantella A, Vickers SM, Banerjee S, Dawra R, Roy S, Ramakrishnan S, Saluja A, Dudeja V (2018) Gut microbiota promotes tumor growth in mice by modulating immune response. Gastroenterology 155(1):33–37 e6. https://doi.org/10.1053/j.gastro.2018.04.001
Shi Z, Park HR, Du Y, Li Z, Cheng K, Sun SY, Li Z, Fu H, Khuri FR (2015) Cables1 complex couples survival signaling to the cell death machinery. Cancer Res 75(1):147–158. https://doi.org/10.1158/0008-5472.CAN-14-0036
Siegel RL, Miller KD (2019) Jemal A (2019) Cancer statistics. CA Cancer J Clin 69(1):7–34. https://doi.org/10.3322/caac.21551
Signoretti M, Roggiolani R, Stornello C, Delle Fave G, Capurso G (2017) Gut microbiota and pancreatic diseases. Minerva Gastroenterol Dietol 63(4):399–410. https://doi.org/10.23736/S1121-421X.17.02387-X
Siolas D, Hannon GJ (2013) Patient-derived tumor xenografts: transforming clinical samples into mouse models. Cancer Res 73(17):5315–5319. https://doi.org/10.1158/0008-5472.CAN-13-1069
Tamanai-Shacoori Z, Smida I, Bousarghin L, Loreal O, Meuric V, Fong SB, Bonnaure-Mallet M, Jolivet-Gougeon A (2017) Roseburia spp.: a marker of health? Future Microbiol 12:157–170. https://doi.org/10.2217/fmb-2016-0130
Tang WH, Wang Z, Levison BS, Koeth RA, Britt EB, Fu X, Wu Y, Hazen SL (2013) Intestinal microbial metabolism of phosphatidylcholine and cardiovascular risk. N Engl J Med 368(17):1575–1584. https://doi.org/10.1056/NEJMoa1109400
van den Munckhof ICL, Kurilshikov A, Ter Horst R, Riksen NP, Joosten LAB, Zhernakova A, Fu J, Keating ST, Netea MG, de Graaf J, Rutten JHW (2018) Role of gut microbiota in chronic low-grade inflammation as potential driver for atherosclerotic cardiovascular disease: a systematic review of human studies. Obes Rev 19(12):1719–1734. https://doi.org/10.1111/obr.12750
Vogelstein B, Papadopoulos N, Velculescu VE, Zhou S, Diaz LA Jr, Kinzler KW (2013) Cancer genome landscapes. Science 339(6127):1546–1558. https://doi.org/10.1126/science.1235122
Wajant H, Pfizenmaier K, Scheurich P (2003) Tumor necrosis factor signaling. Cell Death Differ 10(1):45–65. https://doi.org/10.1038/sj.cdd.4401189
Wan L, Zhou X, Wang C, Chen Z, Peng H, Hou X, Peng Y, Wang P, Li T, Yuan H, Shi Y, Hou X, Xu K, **e Y, He L, **a K, Tang B, Jiang H (2019) Alterations of the gut microbiota in multiple system atrophy patients. Front Neurosci 13:1102. https://doi.org/10.3389/fnins.2019.01102
Wang C, Li J (2015) Pathogenic microorganisms and pancreatic cancer. Gastrointest Tumors 2(1):41–47. https://doi.org/10.1159/000380896
Wei M-Y, Shi S, Liang C, Meng Q-C, Hua J, Zhang Y-Y, Liu J, Zhang B, Xu J, Yu X-J (2019) The microbiota and microbiome in pancreatic cancer: more influential than expected. Mol Cancer 18(1):97. https://doi.org/10.1186/s12943-019-1008-0
Wolfgang CL, Herman JM, Laheru DA, Klein AP, Erdek MA, Fishman EK, Hruban RH (2013) Recent progress in pancreatic cancer. CA Cancer J Clin 63(5):318–348. https://doi.org/10.3322/caac.21190
Wu J, Cai J (2021) Dilemma and challenge of immunotherapy for pancreatic cancer. Dig Dis Sci 66(2):359–368. https://doi.org/10.1007/s10620-020-06183-9
Wu HJ, Ivanov II, Darce J, Hattori K, Shima T, Umesaki Y, Littman DR, Benoist C, Mathis D (2010) Gut-residing segmented filamentous bacteria drive autoimmune arthritis via T helper 17 cells. Immunity 32(6):815–827. https://doi.org/10.1016/j.immuni.2010.06.001
Yang Y, Jobin C (2017) Novel insights into microbiome in colitis and colorectal cancer. Curr Opin Gastroenterol 33(6):422–427. https://doi.org/10.1097/MOG.0000000000000399
Yeste-Velasco M, Mao X, Grose R, Kudahetti SC, Lin D, Marzec J, Vasiljevic N, Chaplin T, Xue L, Xu M, Foster JM, Karnam SS, James SY, Chioni AM, Gould D, Lorincz AT, Oliver RT, Chelala C, Thomas GM, Shipley JM, Mather SJ, Berney DM, Young BD, Lu YJ (2014) Identification of ZDHHC14 as a novel human tumour suppressor gene. J Pathol 232(5):566–577. https://doi.org/10.1002/path.4327
Zhang HL, Yu LX, Yang W, Tang L, Lin Y, Wu H, Zhai B, Tan YX, Shan L, Liu Q, Chen HY, Dai RY, Qiu BJ, He YQ, Wang C, Zheng LY, Li YQ, Wu FQ, Li Z, Yan HX, Wang HY (2012) Profound impact of gut homeostasis on chemically-induced pro-tumorigenic inflammation and hepatocarcinogenesis in rats. J Hepatol 57(4):803–812. https://doi.org/10.1016/j.jhep.2012.06.011
Zheng Y, Wu C, Yang J, Zhao Y, Jia H, Xue M, Xu D, Yang F, Fu D, Wang C, Hu B, Zhang Z, Li T, Yan S, Wang X, Nelson PJ, Bruns C, Qin L, Dong Q (2020) Insulin-like growth factor 1-induced enolase 2 deacetylation by HDAC3 promotes metastasis of pancreatic cancer. Signal Transduct Target Ther 5(1):53. https://doi.org/10.1038/s41392-020-0146-6
Funding
This research was supported by Basic Science Research Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Education (Grant number: 2021R1A2C2005472) and a grant from the Asan Institute for Life Sciences (Grant No. 2020IP0036-1).
Author information
Authors and Affiliations
Contributions
KL and HJO designed the study. KL and M-SK collected the data, performed the main experiments, and wrote the manuscript. SK, SA, and MJK helped with the experiments. SC and S-WK supervised the research progress, led this study, and edited the manuscript. All authors reviewed and approved the manuscript.
Corresponding authors
Ethics declarations
Ethics approval (for animal study)
For the generation of PDX model of pancreatic cancer, procedure for animal experiment was inspected and approved by the Institutional Animal Care and Use Committees (IACUC) of the Asan Institute for Life Sciences (AILS, Project Number: 2019–14-367).
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Lee, K., Oh, H.J., Kang, MS. et al. Metagenomic analysis of gut microbiome reveals a dynamic change in Alistipes onderdonkii in the preclinical model of pancreatic cancer, suppressing its proliferation. Appl Microbiol Biotechnol 105, 8343–8358 (2021). https://doi.org/10.1007/s00253-021-11617-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-021-11617-z