Abstract
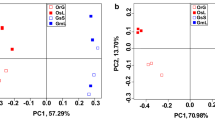
Plants modulate the soil microbiota and select a specific microbial community in the rhizosphere. However, plant domestication reduces genetic diversity, changes plant physiology, and could have an impact on the associated microbiome assembly. Here, we used 16S rRNA gene sequencing to assess the microbial community in the bulk soil and rhizosphere of wild, semi-domesticated, and domesticated genotypes of lima bean (Phaseolus lunatus), to investigate the effect of plant domestication on microbial community assembly. In general, rhizosphere communities were more diverse than bulk soil, but no differences were found among genotypes. Our results showed that the microbial community’s structure was different from wild and semi-domesticated as compared to domesticated genotypes. The community similarity decreased 57.67% from wild to domesticated genotypes. In general, the most abundant phyla were Actinobacteria (21.9%), Proteobacteria (20.7%), Acidobacteria (14%), and Firmicutes (9.7%). Comparing the different genotypes, the analysis showed that Firmicutes (Bacillus) was abundant in the rhizosphere of the wild genotypes, while Acidobacteria dominated semi-domesticated plants, and Proteobacteria (including rhizobia) was enriched in domesticated P. lunatus rhizosphere. The domestication process also affected the microbial community network, in which the complexity of connections decreased from wild to domesticated genotypes in the rhizosphere. Together, our work showed that the domestication of P. lunatus shaped rhizosphere microbial communities from taxonomic to a functional level, changing the abundance of specific microbial groups and decreasing the complexity of interactions among them.






Similar content being viewed by others
Data Availability
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Abdelfattah A, Tack AJM, Wasserman B et al (2021) Evidence for host–microbiome co‐evolution in apple. New Phytol.https://doi.org/10.1111/NPH.17820
Abdullaeva Y, Manirajan BA, Honermeier B, Schnell S, Cardinale M (2021) Domestication affects the composition, diversity, and co-occurrence of the cereal seed microbiota. J. Adv. Res. 31:75–86. https://doi.org/10.1016/j.jare.2020.12.008
Abdullaeva Y, Ratering S, Manirajan BA, Rosado-Porto D, Schnell S, Cardinale M (2022) Domestication impacts the wheat-associated microbiota and the rhizosphere colonization by seed- and soil-originated microbiomes, across different fields. Front. Plant Sci. 12:806915. https://doi.org/10.3389/fpls.2021.806915
Aguilar OM, Riva O, Peltzer E (2004) Analysis of Rhizobium etli and of its symbiosis with wild Phaseolus vulgaris supports coevolution in centers of host diversification. Proc. Natl. Acad. Sci. 101:13548–13553. https://doi.org/10.1073/pnas.0405321101
Anderson MJ (2001) A new method for non parametric multivariate analysis of variance. Austral. Ecol. 26:32–46
Andueza-Noh RH, Martha L, Serrano-Serrano MI et al (2013) Multiple domestications of the Mesoamerican gene pool of lima bean (Phaseolus lunatus L.): evidence from chloroplast DNA sequences. Gen. Res. Crop. Evol. 60:1069–1086
Araujo FF, Bonifacio A, Bavaresco LG et al (2021) Bacillus subtilis changes the root architecture of soybean grown on nutrient-poor substrate. Rhizosphere 18:16–19. https://doi.org/10.1016/j.rhisph.2021.100348
Assunção IP, Nascimento LD, Ferreira MF et al (2011) Reaction of faba bean genotypes to Rhizoctonia solani and resistance stability. Hortic. Bras. 29:492–497. https://doi.org/10.1590/S0102-05362011000400008
Bastian M, Heymann S, Jacomy M (2009) Gephi: An open source software for exploring and manipulating networks. In Proceedings of the Third International ICWSM Conference. California, USA. 361– 362
Bellucci E, Bitocchi E, Ferrarini A et al (2014) Decreased nucleotide and expression diversity and modified coexpression patterns characterize domestication in the common bean. Plant Cell 26:1901–1912. https://doi.org/10.1105/TPC.114.124040
Bitocchi E, Rau D, Bellucci E, Rodriguez M et al (2017) Beans (Phaseolus ssp.) as a model for understanding crop evolution. Front. Pl. Sci. 8:722. https://doi.org/10.3389/fpls.2017.00722
Brisson VL, Schmidt JE, Northen TR (2019) Impacts of maize domestication and breeding on rhizosphere microbial community recruitment from a nutrient depleted agricultural soil. Sci Rep 9:15611. https://doi.org/10.1038/s41598-019-52148-y
Bulgarelli D, Garrido-Oter R, Münch PC et al (2015) Structure and function of the bacterial root microbiota in wild and domesticated barley. Cell Host Microb 17:392–403
Chacón-Sánchez MI, Martínez-Castillo J (2017) Testing domestication scenarios of lima bean (Phaseolus lunatus L.) in Mesoamerica: insights from genome-wide genetic markers. Front. Plant Sci. 8:1551. https://doi.org/10.3389/fpls.2017.01551.
Chouhan GK, Verma JP, Jaiswal DK et al (2021) Phytomicrobiome for promoting sustainable agriculture and food security: Opportunities, challenges, and solutions. Microbiol Res 248:126763
Costa CN, Antunes JEL, Lopes ACA, Freitas ADS, Araujo ASF (2020) Inoculation of rhizobia increases lima bean (Phaseolus lunatus) yield in soils from Piauí and Ceará states, Brazil. Rev Ceres 67:419–423
Doebley JF, Gaut BS, Smith BD (2006) The molecular genetics of crop domestication. Cell 127:1309–1321
Ferguson SJ, Richardson DJ, Van Spanning RJM (2007) Biochemistry and molecular biology of nitrification. Biol. Nitrogen Cycle 209–222https://doi.org/10.1016/B978-044452857-5.50015-1
Fernie AR, Yan J (2019) De Novo domestication: an alternative route toward new crops for the future. Mol. Plant 12:615–631
Freytag GF, Debouck DG (2002) Taxonomy, distribution, and ecology of the genus Phaseolus (Leguminosae-Papilionoideae) in North America, Mexico and Central America. Bot. Res. Ins. Texas.
Friedman J, Alm EJ (2012) Inferring correlation networks from genomic survey data. PLoS Comput. Biol. 8:e1002687
Goss-Souza D, Mendes LW, Rodrigues JLM, Tsai SM (2019) Ecological processes sha** bulk soil and rhizosphere microbiome assembly in a long-term Amazon Forest-to-agriculture conversion. Microb. Ecol. 79:110–122
Gray DA, Dugar G, Gamba P et al (2019) Extreme slow growth as alternative strategy to survive deep starvation in bacteria. Nat. Commun. 101:1–12. https://doi.org/10.1038/s41467-019-08719-8
Gross BL, Olsen KM (2010) Genetic perspectives on crop domestication. Trends Plant Sci. 15:529–537. https://doi.org/10.1016/j.tplants.2010.05.008
Hammer Ø, Harper DAT, Ryan PD (2001) PAST: Paleontological Statistics Software Package for education and data analysis. Palaeontol. Electron. 4:1–9
Kalam S, Basu A, Ahmad I et al (2020) Recent understanding of soil Acidobacteria and their ecological significance: a critical review. Front. Microbiol. 11:580024. https://doi.org/10.3389/fmicb.2020.580024
Ladygina N, Hedlund K (2010) Plant species influence microbial diversity and carbon allocation in the rhizosphere. Soil Biol. Biochem. 42:162–168. https://doi.org/10.1016/j.soilbio.2009.10.009
López-Mondéjar R, Zühlke D, Becher D et al (2016) Cellulose and hemicellulose decomposition by forest soil bacteria proceeds by the action of structurally variable enzymatic systems. Sci. Rep. 6:1–12. https://doi.org/10.1038/srep25279
Mendes LW, Kuramae EE, Navarrete AA, van Veen JA, Tsai SM (2014) Taxonomical and functional microbial community selection in soybean rhizosphere. ISME J. 8:1577–1587
Mendes LW, Mendes R, Raaijmakers JM, Tsai SM (2018a) Breeding for soil-borne pathogen resistance impacts active rhizosphere microbiome of common bean. ISME J.https://doi.org/10.1038/s41396-018-0234-6
Mendes LW, Raaijmakers JM, Hollander M, Mendes R, Tsai SM (2018) Influence of resistance breeding in common bean on rhizosphere microbiome composition and function. ISME J. 12:212–224
Mendes R, Kruijt M, De Bruijn I et al (2011) Deciphering the rhizosphere microbiome for disease-suppressive bacteria. Science 332:1097–1100
Mitter B, Pfaffenbichler N, Flavell R et al (2017) A new approach to modify plant microbiomes and traits by introducing beneficial bacteria at flowering into progeny seeds. Front. Microbiol. 8:1–10. https://doi.org/10.3389/fmicb.2017.00011
Moroenyane I, Mendes LW, Trembley J, Tripathi B, Yergeau É (2021) Plant compartments and developmental stages modulate the balance between niche-based and neutral processes in soybean microbiome. Microbial. Ecol. 82:416–428
Motta-Aldana JR, Serrano-Serrano ML, Hernández-Torres J et al (2010) Multiple Origins of Lima Bean Landraces in the Americas: Evidence from Chloroplast and Nuclear DNA Polymorphisms. Crop Sci. 50:1773
Oliveiros JC (2007) VENNY. An interactive tool for comparing lists with Venn diagrams. http://bioinfogp.cnb.csic.es/tools/venny/index.html.
Pérez-Jaramillo J, Carrión V, Bosse M et al (2017) Linking rhizosphere microbiome composition of wild and domesticated Phaseolus vulgaris to genotypic and root phenotypic traits. ISME J. 11:2244–2257. https://doi.org/10.1038/ismej.2017.85
Pérez-Jaramillo JE, Mendes R, Raaijmakers JM (2016) Impact of plant domestication on rhizosphere microbiome assembly and functions. Plant Mol. Biol. 90:635–644. https://doi.org/10.1007/S11103-015-0337-7
Pérez-Jaramillo HM, Ramírez CA, Mendes R, Raaijmakers JM, Carrión VJ (2019) Deciphering rhizosphere microbiome assembly of wild and modern common bean (Phaseolus vulgaris) in native and agricultural soils from Colombia. Microbiome 7:114
Pickersgill B (2007) Domestication of plants in the Americas: insights from mendelian and molecular genetics. Ann. Botany 100:925–940. https://doi.org/10.1093/aob/mcm193
Praeg N, Illmer P (2020) Microbial community composition in the rhizosphere of Larix decidua under different light regimes with additional focus on methane cycling microorganisms. Sci. Rep. 10:22324. https://doi.org/10.1038/s41598-020-79143-y
Preece C, Livarda A, Christin PA et al (2017) How did the domestication of Fertile Crescent grain crops increase their yields? Funct. Ecol. 31:387–397. https://doi.org/10.1111/1365-2435.12760
Purugganan MD, Fuller DQ (2009) The nature of selection during plant domestication. Nature 457:843–848. https://doi.org/10.1038/nature07895
Rossmann M, Pérez-Jaramillo JE, Kavamura VN et al (2020) Multitrophic interactions in the rhizosphere microbiome of wheat: from bacteria and fungi to protists. FEMS Microbiol. Ecol. 96:fiaa032.
Sansinenea E (2019) Bacillus spp.: As plant growth-promoting bacteria. Secondary Metabolites of Plant Growth Promoting Rhizomicroorganisms. 225–237. https://doi.org/10.1007/978-981-13-5862-3_11
Segata N, Izard J, Waldron L et al (2011) Metagenomic biomarker discovery and explanation. Genome Biol. 12:R60. https://doi.org/10.1186/gb-2011-12-6-r60
Singh J, Sun M, Cannon SB et al (2021) An accumulation of genetic variation and selection across the disease-related genes during apple domestication. Tree Genet. Genomes 17:1–11. https://doi.org/10.1007/S11295-021-01510-1/FIGURES/5
Soldan R, Fusi M, Cardinale M et al (2021) The effect of plant domestication on host control of the microbiota. Commun. Biol. 4:1–9. https://doi.org/10.1038/s42003-021-02467-6
Spor A, Roucou A, Mounier A (2020) Domestication-driven changes in plant traits associated with changes in the assembly of the rhizosphere microbiota in tetraploid wheat. Sci. Rep. 10:12234. https://doi.org/10.1038/s41598-020-69175-9
Van Elsas JD, Chiurazzi M, Mallon CA et al (2012) Microbial diversity determines the invasion of soil by a bacterial pathogen. Proc. Natl. Acad. Sci. USA 109:1159–1164
Wei Z, Jousset A (2017) Plant breeding goes microbial. Trends Plant Sci. 22:555–558. https://doi.org/10.1016/j.tplants.2017.05.009
Wei Z, Yang T, Friman VP et al (2015) Trophic network architecture of root-associated bacterial communities determines pathogen invasion and plant health. Nat. Commun. 6:8413
**ong W, Song Y, Yang K et al (2020) Rhizosphere protists are key determinants of plant health. Microbiome 8:1–9. https://doi.org/10.1186/s40168-020-00799-9
Yadav A, Borrelli JC, Elshahed MS, Youssef NH (2021) Genomic analysis of family UBA6911 (Group 18 Acidobacteria) expands the metabolic capacities of the phylum and highlights adaptations to terrestrial habitats. Appl. Environ. Microbiol. 87:1–19. https://doi.org/10.1128/AEM.00947-21
Acknowledgements
This study was funded by Conselho Nacional de Desenvolvimento Científico e Tecnologico–CNPq (grant 305069/2018-1). and Coordenação de Aperfeiçoamento de Pessoal de Nível Superior–CAPES (Code 001). The authors thank to the Centro de Genética e Bioinformática (CeGenBio) from the Unit of Research (NPDM/UFC). Josieli Lima da Silva, Sandra Mara Barbosa Rocha, Jadson Emanuel Lopes Antunes, and Veronica Brito Silva thank CAPES for their fellowship. Ademir Sergio Ferreira de Araujo, Vania Maria Maciel Melo, Regina Lucia Ferreira Gomes, and Angela Celis de Almeida Lopes thank CNPq for their fellowship of research. Lucas William Mendes thanks FAPESP for his fellowship.
Funding
Conselho Nacional de Desenvolvimento Científico e Tecnologico – CNPq (grant 305069/2018–1).
Author information
Authors and Affiliations
Contributions
ASFA, ACAL, and RLFG conceived this study. JLS, SMBR, JELA, and LMSO conducted the experiment, collected samples, and proceeded DNA extraction. VMMM and FASO provided the sequencing. APAP and LWM provided the bioinformatic and the 16S rRNA gene data. LWM, APAP, GNC, and VBS performed the statistical, network analyses and generate the results. ASFA, APAP, LWM, VMMM, ACAL, and FAN interpreted the results and elaborated the main arguments. All authors wrote and reviewed and the final manuscript.
Corresponding author
Ethics declarations
Ethics Approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Consent to Participate
Not applicable
Consent for Publication
Not applicable.
Competing Interests
The authors declare no competing interests.
Rights and permissions
About this article
Cite this article
da Silva, J.L., Mendes, L.W., Rocha, S.M.B. et al. Domestication of Lima Bean (Phaseolus lunatus) Changes the Microbial Communities in the Rhizosphere. Microb Ecol 85, 1423–1433 (2023). https://doi.org/10.1007/s00248-022-02028-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-022-02028-2