Abstract
Presenilin proteins make the catalytic component of γ-secretase, a multiprotein transmembrane protease, and are type II transmembrane proteins. Amyloid protein, Notch, and beta catenin are among more than 90 substrates of Presenilins. Mutations in Presenilins lead to defects in proteolytic cleavage of its substrate resulting in some of the most devastating pathological conditions including Alzheimer disease (AD), developmental disorders, and cancer. In addition to catalytic roles, Presenilin protein is also shown to be involved in many non-catalytic roles, i.e., calcium homeostasis, regulation of autophagy, and protein trafficking, etc. These proteolytic proteins are highly conserved and are present in almost all the major eukaryotic groups. Studies, performed on a wide variety of organisms ranging from human to unicellular dictyostelium, have shown the important catalytic and non-catalytic roles of Presenilins. In this study, we infer the evolutionary patterns and history of Presenilins as well as of other γ-secretase proteins. We show that Presenilins are the most ancient of the γ-secretase proteins and that Presenilins may have their origin in the last common ancestor (LCA) of Eukaryotes. We also demonstrate that Presenilin proteins generally lack diversifying selection during the course of their evolution. Through evolutionary trace analysis, we show that Presenilin protein sites that undergo mutations in Familial Alzheimer disease, are highly conserved in metazoans. Finally, we discuss the evolutionary, physiological, and pathological implications of our findings and propose that the evolutionary profile of Presenilins supports the loss of function hypothesis of AD pathogenesis.



taken from the work of Zhang et al. (2013). c Bar graph showing the frequency of rank values/conservation score (with 1 being for the most conserved and 10 being the least conserved residue) with their corresponding rank values/conservation scores in the transmembrane and non-transmembrane portion of the Presenilin-1 protein. d List of site-directed mutations (taken from UniProt database) which have been carried out in the Presenilin-1 gene. Column-1 represents the position of residues; column-2 shows the heatmap, depicting conservation profile of mutated sites; column-3 represents substituted residues on the left side of the arrow and replacement residues on the right side of the arrow with more than one replacement residues giving rise to a specific phenotype are separated by a comma, and finally, the phenotypic effects due to these mutations are given in columns 4 where Φ-Loss of interaction with GFAP, Ω-abolished protease activity, Ψ-increased protease activity, ∆-abolished caspase cleavage, α-alters γ-secretase specificity, Π-abolished PKA signaling, and β-reduced notch processing (Color figure online)


Similar content being viewed by others
References
Albà MM, Castresana J (2005) Inverse relationship between evolutionary rate and age of mammalian genes. Mol Biol Evol 22:598–606. https://doi.org/10.1093/molbev/msi045
Ali R, Muhammad S, Khan M, Arvestad L (2013) Quantitative synteny scoring improves homology inference and partitioning of gene families. BMC Bioinform. https://doi.org/10.1186/1471-2105-14-S15-S12
Ali RH, Muhammad SA, Arvestad L (2016) GenFamClust: an accurate, synteny-aware and reliable homology inference algorithm. BMC Evol Biol. https://doi.org/10.1186/s12862-016-0684-2
Ali RH, Bogusz M, Whelan S, Tamura K (2019) Identifying clusters of high confidence homologies in multiple sequence alignments. Mol Biol Evol. https://doi.org/10.1093/molbev/msz142
Altschul SF, Madden TL, Schäffer AA et al (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402
Area-Gomez E, de Groof AJC, Boldogh I et al (2009) Presenilins are enriched in endoplasmic reticulum membranes associated with mitochondria. Am J Pathol 175:1810–1816. https://doi.org/10.2353/ajpath.2009.090219
Bai XC, Yan C, Yang G et al (2015) An atomic structure of human gamma-secretase. Nature 525:212–217. https://doi.org/10.1038/nature14892
Beel AJ, Sanders CR (2008) Substrate specificity of gamma-secretase and other intramembrane proteases. Cell Mol Life Sci 65:1311–1334. https://doi.org/10.1007/s00018-008-7462-2
Blekhman R, Man O, Herrmann L et al (2008) Natural selection on genes that underlie human disease susceptibility. Curr Biol 18:883–889. https://doi.org/10.1016/j.cub.2008.04.074
Bouckaert R, Vaughan TG, Barido-Sottani J et al (2019) BEAST 2.5: an advanced software platform for Bayesian evolutionary analysis. PLoS Comput Biol. https://doi.org/10.1371/journal.pcbi.1006650
Brown MS, Ye J, Rawson RB, Goldstein JL (2000) Regulated intramembrane proteolysis: a control mechanism conserved from bacteria to humans. Cell 100:391–398. https://doi.org/10.1016/S0092-8674(00)80675-3
Brunkan AL, Martinez M, Walker ES, Goate AM (2005) Presenilin endoproteolysis is an intramolecular cleavage. Mol Cell Neurosci 29:65–73. https://doi.org/10.1016/j.mcn.2004.12.012
Bustos V, Pulina MV, Bispo A et al (2017a) Phosphorylated Presenilin 1 decreases β-amyloid by facilitating autophagosome-lysosome fusion. Proc Natl Acad Sci USA. https://doi.org/10.1073/pnas.1705240114
Bustos V, Pulina MV, Kelahmetoglu Y et al (2017b) Bidirectional regulation of Aβ levels by Presenilin 1. Proc Natl Acad Sci USA. https://doi.org/10.1073/pnas.1705235114
Cai D, Leem JY, Greenfield JP et al (2003) Presenilin-1 regulates intracellular trafficking and cell surface delivery of beta-amyloid precursor protein. J Biol Chem 278:3446–3454. https://doi.org/10.1074/jbc.M209065200
Capell A, Steiner H, Romig H et al (2000) Presenilin-1 differentially facilitates endoproteolysis of the beta-amyloid precursor protein and Notch. Nat Cell Biol 2:205–211. https://doi.org/10.1038/35008626
De Strooper B, Beullens M, Contreras B et al (1997) Phosphorylation, subcellular localization, and membrane orientation of the Alzheimer’s disease-associated presenilins. J Biol Chem 272:3590–3598. https://doi.org/10.1074/jbc.272.6.3590
Degnan JH, Rosenberg NA (2009) Gene tree discordance, phylogenetic inference and the multispecies coalescent. Trends Ecol Evol 24:332–340
DeLuna A, Vetsigian K, Shoresh N et al (2008) Exposing the fitness contribution of duplicated genes. Nat Genet. https://doi.org/10.1038/ng.123
Domazet-Loso T, Tautz D (2003) An evolutionary analysis of orphan genes in Drosophila. Genome Res 13:2213–2219. https://doi.org/10.1101/gr.1311003
Drenos F, Kirkwood TBL (2010) Selection on alleles affecting human longevity and late-life disease: the example of apolipoprotein E. PLoS ONE. https://doi.org/10.1371/journal.pone.0010022
Duff K, Eckman C, Zehr C et al (1996) Increased amyloid-β42(43) in brains of mice expressing mutant presenilin 1. Nature 383:710–713. https://doi.org/10.1038/383710a0
Edbauer D, Winkler E, Regula JT et al (2003) Reconstitution of gamma-secretase activity. Nat Cell Biol 5:486–488. https://doi.org/10.1038/ncb960
Espagne E, Lespinet O, Malagnac F et al (2008) The genome sequence of the model ascomycete fungus Podospora anserina. Genome Biol. https://doi.org/10.1186/gb-2008-9-5-r77
Fan HW, Noda H, **e HQ et al (2015) Genomic analysis of an ascomycete fungus from the rice planthopper reveals how it adapts to an endosymbiotic lifestyle. Genome Biol Evol. https://doi.org/10.1093/gbe/evv169
Fukumori A, Fluhrer R, Steiner H, Haass C (2010) Three-amino acid spacing of presenilin endoproteolysis suggests a general stepwise cleavage of -secretase-mediated intramembrane proteolysis. J Neurosci 30:7853–7862. https://doi.org/10.1523/JNEUROSCI.1443-10.2010
Goutte C, Tsunozaki M, Hale VA, Priess JR (2002) APH-1 is a multipass membrane protein essential for the Notch signaling pathway in Caenorhabditis elegans embryos. Proc Natl Acad Sci USA 99:775–779. https://doi.org/10.1073/pnas.022523499
Gu Z, Steinmetz LM, Gu X et al (2003) Role of duplicate genes in genetic robustness against null mutations. Nature. https://doi.org/10.1038/nature01198
Han MV, Zmasek CM (2009) PhyloXML: XML for evolutionary biology and comparative genomics. BMC Bioinform 10:356. https://doi.org/10.1186/1471-2105-10-356
Hirsh AE, Fraser HB (2001) Protein dispensability and rate of evolution. Nature 411:1046–1049. https://doi.org/10.1038/35082561
Jacobsen H, Reinhardt D, Brockhaus M et al (1999) The influence of endoproteolytic processing of familial Alzheimer’s disease presenilin 2 on Aβ42 amyloid peptide formation. J Biol Chem 274:35233–35239. https://doi.org/10.1074/jbc.274.49.35233
Juretić D, Zoranić L, Zucić D (2002) Basic charge clusters and predictions of membrane protein topology. J Chem Inf Comput Sci. https://doi.org/10.1021/ci010263s
Kang DE, Soriano S, **a X et al (2002) Presenilin couples the paired phosphorylation of β-catenin independent of axin: Implications for β-catenin activation in tumorigenesis. Cell. https://doi.org/10.1016/S0092-8674(02)00970-4
Keeling PJ, Slamovits CH (2005) Causes and effects of nuclear genome reduction. Curr Opin Genet Dev 15:601
Kelleher RJ, Shen J (2017) Presenilin-1 mutations and Alzheimer’s disease. Proc Natl Acad Sci USA 114:629–631. https://doi.org/10.1073/pnas.1619574114
Kirkwood TBL, Austad SN (2000) Why do we age? Nature 408:233–238. https://doi.org/10.1038/35041682
Koonin EV (2005) Orthologs, paralogs, and evolutionary genomics. Annu Rev Genet. https://doi.org/10.1146/annurev.genet.39.073003.114725
Kosakovsky Pond SL, Frost SDW (2005) Datamonkey: rapid detection of selective pressure on individual sites of codon alignments. Bioinformatics 21:2531–2533. https://doi.org/10.1093/bioinformatics/bti320
Kulic L, Walter J, Multhaup G et al (2000) Separation of presenilin function in amyloid beta-peptide generation and endoproteolysis of Notch. Proc Natl Acad Sci USA 97:5913–5918. https://doi.org/10.1073/pnas.100049897
Kumar S, Stecher G, Suleski M, Hedges SB (2017) TimeTree: a resource for timelines, timetrees, and divergence times. Mol Biol Evol. https://doi.org/10.1093/molbev/msx116
LaFerla FM (2002) Calcium dyshomeostasis and intracellular signalling in alzheimer’s disease. Nat Rev Neurosci 3:862–872. https://doi.org/10.1038/nrn960
Laudon H, Hansson EM, Melén K et al (2005) A nine-transmembrane domain topology for presenilin 1. J Biol Chem 280:35352–35360. https://doi.org/10.1074/jbc.M507217200
Lee MK, Slunt HH, Martin LJ et al (1996) Expression of presenilin 1 and 2 (PS1 and PS2) in human and murine tissues. J Neurosci 16:7513–7525
Levy-Lahad E, Wasco W, Poorkaj P et al (1995) Candidate gene for the chromosome 1 familial Alzheimer’s disease locus. Science 269:973–977
Lichtarge O, Bourne HR, Cohen FE (1996) An evolutionary trace method defines binding surfaces common to protein families. J Mol Biol 257:342–358. https://doi.org/10.1006/jmbi.1996.0167
Lua RC, Lichtarge O (2010) PyETV: a PyMOL evolutionary trace viewer to analyze functional site predictions in protein complexes. Bioinformatics 26:2981–2982. https://doi.org/10.1093/bioinformatics/btq566
Ludtmann MHR, Otto GP, Schilde C et al (2014) An ancestral non-proteolytic role for presenilin proteins in multicellular development of the social amoeba dictyostelium discoideum. J Cell Sci. https://doi.org/10.1242/jcs.140939
Maddison WP (1997) Gene trees in species trees. Syst Biol. https://doi.org/10.1093/sysbio/46.3.523
Marchler-Bauer A, Bo Y, Han L et al (2016) CDD/SPARCLE: Functional classification of proteins via subfamily domain architectures. Nucleic Acids Res. https://doi.org/10.1093/nar/gkw1129
McCarthy JV, Twomey C, Wujek P (2009) Presenilin-dependent regulated intramembrane proteolysis and γ-secretase activity. Cell Mol Life Sci 66:1534–1555
Menzies FM, Fleming A, Rubinsztein DC (2015) Compromised autophagy and neurodegenerative diseases. Nat Rev Neurosci 16:345–357. https://doi.org/10.1038/nrn3961
Meyer DZ, Avery LM (2009) Excel as a qualitative data analysis tool. Field Methods 21:91–112. https://doi.org/10.1177/1525822X08323985
Murray MM, Jones CJ, Clark RF (2000) The presenilin gene is very highly conserved throughout evolution. Neurobiol Aging 21:48. https://doi.org/10.1016/S0197-4580(00)82893-6
Murrell B, Wertheim JO, Moola S et al (2012) Detecting individual sites subject to episodic diversifying selection. PLoS Genet. https://doi.org/10.1371/journal.pgen.1002764
Nixon RA, Yang DS (2011) Autophagy failure in Alzheimer’s disease-locating the primary defect. Neurobiol Dis 43:38–45
Ohno S (1970) Evolution by gene duplication. Springer, Berlin
Ohta K, Mizuno A, Ueda M et al (2010) Autophagy impairment stimulates PS1 expression and γ-secretase activity. Autophagy 6:345–352
Otto GP, Sharma D, Williams RSB (2016) Non-catalytic roles of presenilin throughout evolution. J Alzheimer’s Dis 52:1177–1187
Pettersen EF, Goddard TD, Huang CC et al (2004) UCSF chimera—a visualization system for exploratory research and analysis. J Comput Chem 25:1605–1612. https://doi.org/10.1002/jcc.20084
Podlisny MB, Citron M, Amarante P et al (1997) Presenilin proteins undergo heterogeneous endoproteolysis between Thr291and Ala299and occur as stable N- and C-terminal fragments in normal and alzheimer brain tissue. Neurobiol Dis 3:325–337. https://doi.org/10.1006/nbdi.1997.0129
Ponting CP (2002) Identification of a novel family of presenilin homologues. Hum Mol Genet. https://doi.org/10.1093/hmg/11.9.1037
Popugaeva E, Bezprozvanny I (2013) Role of endoplasmic reticulum Ca2+ signaling in the pathogenesis of Alzheimer disease. Front Mol Neurosci. https://doi.org/10.3389/fnmol.2013.00029
Pundir S, Martin MJ, O’Donovan C (2017) UniProt protein knowledgebase. In: Wu CH, Arighi CN, Ross KE (eds) Methods in molecular biology. Humana Press, New York, pp 41–55
Sakharkar KR, Chow VTK (2005) Strategies for genome reduction in microbial genomes. Genome Inform 16:69–75
Sato C, Morohashi Y, Tomita T, Iwatsubo T (2006) Structure of the catalytic pore of gamma-secretase probed by the accessibility of substituted cysteines. J Neurosci 26:12081–12088. https://doi.org/10.1523/JNEUROSCI.3614-06.2006
Sato C, Takagi S, Tomita T, Iwatsubo T (2008) The C-terminal PAL motif and transmembrane domain 9 of presenilin 1 are involved in the formation of the catalytic pore of the gamma-secretase. J Neurosci 28:6264–6271. https://doi.org/10.1523/JNEUROSCI.1163-08.2008
Scheper W, Zwart R, Baas F (2004) Rab6 membrane association is dependent of Presenilin 1 and cellular phosphorylation events. Mol Brain Res 122:17–23. https://doi.org/10.1016/j.molbrainres.2003.11.013
Shen J, Kelleher RJ (2007) The presenilin hypothesis of Alzheimer’s disease: evidence for a loss-of-function pathogenic mechanism. Proc Natl Acad Sci USA 104:403–409. https://doi.org/10.1073/pnas.0608332104
Sievers F, Higgins DG (2014a) Clustal omega. Curr Protoc Bioinform. https://doi.org/10.1002/0471250953.bi0313s48
Sievers F, Higgins DG (2014b) Clustal omega, accurate alignment of very large numbers of sequences. Methods Mol Biol. https://doi.org/10.1007/978-1-62703-646-7_6
Slamovits CH (2013) Extreme genome reduction in microbial parasites
Spasic D, Tolia A, Dillen K et al (2006) Presenilin-1 maintains a nine-transmembrane topology throughout the secretory pathway. J Biol Chem 281:26569–26577. https://doi.org/10.1074/jbc.M600592200
Suga K, Tomiyama T, Mori H, Akagawa K (2004) Syntaxin 5 interacts with presenilin holoproteins, but not with their N- or C-terminal fragments, and affects beta-amyloid peptide production. Biochem J 381:619–628. https://doi.org/10.1042/BJ20040618
Sun L, Zhou R, Yang G, Shi Y (2017) Analysis of 138 pathogenic mutations in presenilin-1 on the in vitro production of Aβ42 and Aβ40 peptides by γ-secretase. Proc Natl Acad Sci USA 114:E476–E485. https://doi.org/10.1073/pnas.1618657114
Supnet C, Bezprozvanny I (2010) The dysregulation of intracellular calcium in Alzheimer disease. Cell Calcium 47:183–189
Tajima F (1993) Simple methods for testing the molecular evolutionary clock hypothesis. Genetics 135:599–607
Takasugi N, Tomita T, Hayashi I et al (2003) The role of presenilin cofactors in the γ-secretase complex. Nature 422:438–441. https://doi.org/10.1038/nature01506
Tamura K, Stecher G, Peterson D et al (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729. https://doi.org/10.1093/molbev/mst197
Tharp WG, Sarkar IN (2013) Origins of amyloid-β. BMC Genomics. https://doi.org/10.1186/1471-2164-14-290
Thinakaran G, Borchelt DR, Lee MK et al (1996) Endoproteolysis of presenilin 1 and accumulation of processed derivatives in vivo. Neuron 17:181–190. https://doi.org/10.1016/S0896-6273(00)80291-3
Tolia A, Chávez-Gutiérrez L, De Strooper B (2006) Contribution of presenilin transmembrane domains 6 and 7 to a water-containing cavity in the ??-secretase complex. J Biol Chem 281:27633–27642. https://doi.org/10.1074/jbc.M604997200
Tolia A, Horré K, De Strooper B (2008) Transmembrane domain 9 of presenilin determines the dynamic conformation of the catalytic site of γ-secretase. J Biol Chem 283:19793–19803. https://doi.org/10.1074/jbc.M802461200
Walter J, Capell A, Grünberg J et al (1996) The Alzheimer’s disease-associated presenilins are differentially phosphorylated proteins located predominantly within the endoplasmic reticulum. Mol Med 2:673–691
Wang J, Beher D, Nyborg AC et al (2006a) C-terminal PAL motif of presenilin and presenilin homologues required for normal active site conformation. J Neurochem 96:218–227. https://doi.org/10.1111/j.1471-4159.2005.03548.x
Wang R, Tang P, Wang P et al (2006b) Regulation of tyrosinase trafficking and processing by presenilins: partial loss of function by familial Alzheimer’s disease mutation. Proc Natl Acad Sci USA 103:353–358. https://doi.org/10.1073/pnas.0509822102
Watanabe H, Shen J (2017) Dominant negative mechanism of Presenilin-1 mutations in FAD. Proc Natl Acad Sci USA. https://doi.org/10.1073/pnas.1717180114
Watanabe N, Takagi S, Tominaga A et al (2010) Functional analysis of the transmembrane domains of presenilin 1: Participation of transmembrane domains 2 and 6 in the formation of initial substrate-binding site of γ-secretase. J Biol Chem 285:19738–19746. https://doi.org/10.1074/jbc.M110.101287
Waterhouse AM, Procter JB, Martin DMA et al (2009) Jalview Version 2-A multiple sequence alignment editor and analysis workbench. Bioinformatics. https://doi.org/10.1093/bioinformatics/btp033
Webb B, Sali A (2016) Comparative protein structure modeling using MODELLER. Curr Protoc Bioinform. https://doi.org/10.1002/cpbi.3
Wilkins A, Erdin S, Lua R, Lichtarge O (2012) Evolutionary trace for prediction and redesign of protein functional sites. Methods Mol Biol. https://doi.org/10.1007/978-1-61779-465-0_3
Wilson AC, Carlson SS, White TJ (1977) Biochemical evolution. Annu Rev Biochem 46:573–639. https://doi.org/10.1146/annurev.bi.46.070177.003041
Wolf Y, Novichkov P, Karev G et al (2009) The universal distribution of evolutionary rates of genes and distinct characteristics of eukaryotic genes of different apparent ages. Proc Natl Acad Sci USA 106:7273–7280. https://doi.org/10.1073/pnas.0901808106
Zhang Z, Nadeau P, Song W et al (2000) Presenilins are required for γ-secretase cleavage of β-APP and transmembrane cleavage of Notch-1. Nat Cell Biol. https://doi.org/10.1038/35017108
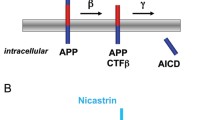
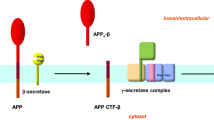
Zhang Y, Thompson R, Zhang H, Xu H (2011) APP processing in Alzheimer’s disease. Mol Brain 4:3. https://doi.org/10.1186/1756-6606-4-3
Acknowledgements
We are grateful to Professor Dr. Khalid J. Siddiqui for spending time in improving the grammar and other language-related aspects of our manuscript.
Author information
Authors and Affiliations
Corresponding author
Additional information
Handling editor: Joana Projecto-Garcia.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Khan, A.A., Ali, R.H. & Mirza, B. Evolutionary History of Alzheimer Disease-Causing Protein Family Presenilins with Pathological Implications. J Mol Evol 88, 674–688 (2020). https://doi.org/10.1007/s00239-020-09966-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-020-09966-w