Abstract
Imaging mass spectrometry (IMS) is a technique in full expansion used in many clinical and biological applications. A common limitation of the technology, particularly true for protein analysis, is that only the most abundant and/or more easily ionizable molecules are typically detected. One approach to overcome this limitation is to transfer proteins contained within tissue sections onto functionalized surfaces with high spatial fidelity for IMS analysis. In this case, only proteins with an affinity for the surface will be retained whereas others will be removed. The chemical nature of the surface is therefore critical. The research work presented herein proposes a high spatial fidelity transfer method for proteins from thin tissue sections onto a nitrocellulose surface. The method employs a home-built apparatus that allows the transfer process to be performed without any direct physical contact between the section and the transfer surface while maintaining physical pressure between the surfaces to help protein migration. The performance of this system was demonstrated using mouse liver and kidney sections. Serials sections were also collected either to be stained with hematoxylin and eosin (H&E) to assess the spatial fidelity of the transfer process or to be directly analyzed as a control sample to differentiate the signals detected after transfer. IMS results showed a high spatial fidelity transfer of a subset of proteins. Some of the detected proteins were poorly observed or not observed with conventional direct tissue analysis, demonstrating an increase in detection sensitivity and specificity with the newly developed method.

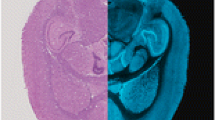
Imaging MS of proteins transferred from tissue sections to a capture membrane





Similar content being viewed by others
References
Cornett DS, Reyzer ML, Chaurand P, Caprioli RM (2007) MALDI imaging mass spectrometry: molecular snapshots of biochemical systems. Nat Meth 4(10):828–833
Norris JL, Caprioli RM (2013) Analysis of tissue specimens by matrix-assisted laser desorption/ionization imaging mass spectrometry in biological and clinical research. Chem Rev 113(4):2309–2342
Stoeckli M, Chaurand P, Hallahan DE, Caprioli RM (2001) Imaging mass spectrometry: a new technology for the analysis of protein expression in mammalian tissues. Nat Med 7:493–496
Chughta K, Heeren RMA (2010) Mass spectrometric imaging for biomedical tissue analysis. Chem Rev 110:3237–3277
Thomas A, Chaurand P (2014) Advances in tissue section preparation for MALDI imaging MS. Bioanalysis 6(7):967–982
Chaurand P, Fouchecourt S, Dague BB, Xu BJ, Reyzer ML, Orgebin-Crist M-C, Caprioli RM (2003) Profiling and imaging proteins in the mouse epididymis by imaging mass spectrometry. Proteomics 3:2221–2239
Chaurand P, Rahman MA, Hunt T, Mobley JA, Gu G, Lathan JC, Caprioli RM, Kasper S (2008) Monitoring mouse prostate development by profiling and imaging mass spectrometry. Mol Cell Proteomics 7:411–423
Chaurand P, Sanders ME, Jensen RA, Caprioli RM (2004) Proteomics in diagnostic pathology: profiling and imaging proteins directly in tissue sections. Am J Pathol 165:1057–1068
Norris JL, Caprioli RM (2013) Imaging mass spectrometry: a new tool for pathology in a molecular age. Proteomics Clin Appl 7(11–12):733–738
Schwamborn K, Caprioli RM (2010) MALDI imaging mass spectrometry—painting molecular pictures. Mol Oncol 4(6):529–538
Reyzer ML, Caldwell RL, Dugger TC, Forbes JT, Ritter CA, Guix M, Arteaga CL, Caprioli RM (2004) Early changes in protein expression detected by mass spectrometry predict tumor response to molecular therapeutics. Cancer Res 64(24)
Reyzer ML, Caprioli RM (2007) MALDI-MS-based imaging of small molecules and proteins in tissues. Curr Opin Chem Biol 11(1):29–35
Chaurand P, Norris JL, Cornett DS, Mobley JA, Caprioli RM (2006) New developments in profiling and imaging of proteins from tissue sections by MALDI mass spectrometry. J Proteome Res 5:2889–2900
Thomas A, Laveaux Charbonneau J, Fournaise E, Chaurand P (2012) Sublimation of new matrix candidates for high spatial resolution imaging mass spectrometry of lipids: enhanced information in both positive and negative polarities after 1,5-diaminonaphthalene deposition. Anal Chem 84:2048–2054
Dufresne M, Thomas A, Breault-Turcot J, Masson J-F, Chaurand P (2013) Silver assisted laser desorption ionization for high spatial resolution imaging mass spectrometry of olefins from thin tissue sections. Anal Chem 85:3318–3324
Chaurand P, Caprioli RM (2002) Direct profiling and imaging of peptides and proteins from mammalian cells and tissue sections by mass spectrometry. Electrophoresis 23(18):3125–3135
Liu R, Li Q, Smith LM (2014) Detection of large ions in time-of-flight mass spectrometry: effects of ion mass and acceleration voltage on microchannel plate detector response. J Am Soc Mass Spectrom 25(8):1374–1383
Lemaire R, Stauber J, Wisztorski M, Van Camp C, Desmons A, Deschamps M, Proess G, Rudlof I, Woods AS, Day R, Salzet M, Fournier I (2007) Tag-mass: specific molecular imaging of transcriptome and proteome by mass spectrometry based on photocleavable tag. J Proteome Res 6(6):2057–2067
Thiery G, Anselmi E, Audebourg A, Darii E, Abarbri M, Terris B, Tabet J-C, Gut IG (2008) Improvements of TArgeted multiplex mass spectrometry IMaging. Proteomics 8(18):3725–3734
Thiery G, Shchepinov MS, Southern EM, Audebourg A, Audard V, Terris B, Gut IG (2007) Multiplex target protein imaging in tissue sections by mass spectrometry—TAMSIM. Rapid Commun Mass Spectrom 21(6):823–829
Yang J, Chaurand P, Norris JL, Porter NA, Caprioli RM (2012) Activity-based probes linked with laser-cleavable mass tags for signal amplification in imaging mass spectrometry: analysis of serine hydrolase enzymes in mammalian tissue. Anal Chem 84(8):3689–3695
Caprioli RM, Farmer TB, Gile J (1997) Molecular imaging of biological samples: localization of peptides and proteins using MALDI-TOF MS. Anal Chem 69(23):4751–4760
Chaurand P, Dague BB, Pearsall RS, Threadgill DW, Caprioli RM (2001) Profiling proteins from azoxymethane-induced colon tumors at the molecular level by MALDI mass spectrometry. Proteomics 1:1320–1326
Chaurand P, Stoeckli M, Caprioli RM (1999) Direct profiling of proteins in biological tissue sections by maldi mass spectrometry. Anal Chem 71:5263–5270
Bouamrani A, Ternier J, Ratel D, Benabid A-L, Issartel J-P, Brambilla E, Berger F (2006) Direct-tissue SELDI-TOF mass spectrometry analysis: a new application for clinical proteomics. Clin Chem 52:2103–2105
Towbin H, Staehelin T, Gordon J (1979) Electrophoretic transfer of proteins from polyacrylamide gels to nitrocellulose sheets—procedure and some applications. Proc Natl Acad Sci U S A 76(9):4350–4354
Luque-Garcia JL, Zhou G, Sun T-T, Neubert TA (2006) Use of nitrocellulose membranes for protein characterization by matrix-assisted laser desorption/ionization mass spectrometry. Anal Chem 78(14):5102–5108
Lobert S, Correia JJ (1994) Method for rapid electrophoretic transfer of isoelectric-focusing gels to polyvinylidene difluoride. Electrophoresis 15(7):930–931
Chaurand P, Cornett DS, Caprioli RM (2006) Molecular imaging of thin mammalian tissue sections by mass spectrometry. Curr Opin Biotechnol 17(4):431–436
Tang N, Tornatore P, Weinberger SR (2004) Current developments in SELDI affinity technology. Mass Spectrom Rev 23:34–44
Müller M, Gras R, Appel RD, Bienvenut WV, Hochstrasser DF (2002) Visualization and analysis of molecular scanner peptide mass spectra. J Am Soc Mass Spectrom 13(3):221–231
Acknowledgments
The authors would like to thank Maxime Couture and Jean-Francois Masson (Department of Chemistry, University of Montreal) for their help with the AFM measurements, as well as Jean-Francois Myre and Louis Beaumont (Department of Chemistry, University of Montreal) for their help and useful insights in the design and building of the transfer and spin coater apparatus. The authors also acknowledge financial support from the Fonds de recherche du Québec—Nature et technologies, the Natural Sciences and Engineering Research Council of Canada, and the Canadian Foundation for Innovation.
Author information
Authors and Affiliations
Corresponding author
Additional information
Published in the topical collection Mass Spectrometry Imaging with guest editors Andreas Römpp and Uwe Karst.
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 8951 kb)
Rights and permissions
About this article
Cite this article
Fournaise, E., Chaurand, P. Increasing specificity in imaging mass spectrometry: high spatial fidelity transfer of proteins from tissue sections to functionalized surfaces. Anal Bioanal Chem 407, 2159–2166 (2015). https://doi.org/10.1007/s00216-014-8300-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-014-8300-z