Abstract
Key message
The Mcgy1 locus responsible for gynoecy was fine-mapped into a 296.94-kb region, in which four single-nucleotide variations and six genes adjacent to them might be associate with sex differentiation in bitter gourd.
Abstract
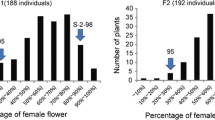
Gynoecy plays an important role in high-efficiency hybrid seed production, and gynoecious plants are excellent materials for dissecting sex differentiation in Cucurbitaceae crop species, including bitter gourd. However, the gene responsible for gynoecy in bitter gourd is unknown. Here, we first identified a gynoecy locus designated Mcgy1 using the F2 population (n = 291) crossed from the gynoecious line S156G and the monoecious line K8-201 via bulked segregant analysis with whole-genome resequencing (BSA-seq) and molecular marker linkage analysis. Then, a large S156G × K8-201 F2 population (n = 5,656) was used for fine-map** to delimit the Mcgy1 locus into a 296.94-kb physical region on pseudochromosome MC01, where included 33 annotated genes different from any homologous gynoecy genes previously reported in Cucurbitaceae species. Within this region, four underlying single-nucleotide variations (SNVs) that might cause gynoecy were identified by multiple genomic sequence variation analysis, and their six neighbouring genes were considered as potential candidate genes for Mcgy1. Of these, only MC01g1681 showed a significant differential expression at two-leaf developmental stage between S156G and its monoecious near-isogenic line S156 based on RNA sequencing (RNA-seq) and qRT-PCR analyses. In addition, transcriptome analysis revealed 21 key differentially expressed genes (DEGs) and possible regulatory pathways of the formation of gynoecy in bitter gourd. Our findings provide a new clue for researching on gynoecious plants in Cucurbitaceae species and a theoretical basis for breeding gynoecious bitter gourd lines by the use of molecular markers-assisted selection.






Similar content being viewed by others
Data availability
The data that support the findings of the current study are available from the corresponding author upon reasonable request. Raw data of genome and transcriptome generated in this study have been deposited in National Genomic Data Center (NGDC) of China National Center for Bioinformation (CNCB) under accession number PRJCA011401.
References
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410. https://doi.org/10.1016/S0022-2836(05)80360-2
An JJ, Gharami K, Liao GY, Woo NH, Lau AG, Vanevski F, Torre ER, Jones KR, Feng Y, Lu B, Xu B (2008) Distinct role of long 3′ UTR BDNF mRNA in spine morphology and synaptic plasticity in hippocampal neurons. Cell 134:175–187. https://doi.org/10.1016/j.cell.2008.05.045
Barrett SCH (2002) The evolution of plant sexual diversity. Nat Rev Genet 3:274–284. https://doi.org/10.1038/nrg776
Barry RM, Bitbol AF, Lorestani A, Charles EJ, Habrian CH, Hansen JM, Li HJ, Baldwin EP, Wingreen NS, Kollman JM, Gitai Z (2014) Large-scale filament formation inhibits the activity of CTP synthetase. Elife 3:e03638. https://doi.org/10.7554/eLife.03638
Basch E, Gabardi S, Ulbricht C (2003) Bitter melon (Momordica charantia): a review of efficacy and safety. Am J Health Syst Pharm 60:356–359. https://doi.org/10.1093/ajhp/60.4.356
Behera TK, Dey SS, Munshi AD, Gaikwad AB, Pal A, Singh I (2009) Sex inheritance and development of gynoecious hybrids in bitter gourd (Momordica charantia L.). Sci Hortic 120:130–133. https://doi.org/10.1016/j.scienta.2008.09.006
Behera TK, Dey SS, Sirohi PS (2006) DBGy-201 and DBGy-202: two gynoecious lines in bitter gourd (Momordica charantia L.) isolated from indigenous source. Indian J Genet Pl Br 66:61–62
Berkovits BD, Mayr C (2015) Alternative 3′ UTRs act as scaffolds to regulate membrane protein localization. Nature 522:363–367. https://doi.org/10.1038/nature14321
Beyer EJ (1976) Silver ion: a potent antiethylene agent in cucumber and tomato. HortScience 11: 195–196. https://journals.ashs.org/hortsci/view/journals/hortsci/11/3/article-p195.xml
Boualem A, Fergany M, Fernandez R, Troadec C, Martin A, Morin H, Sari MA, Collin F, Flowers JM, Pitrat M, Purugganan MD, Dogimont C, Bendahmane A (2008) A conserved mutation in an ethylene biosynthesis enzyme leads to andromonoecy in melons. Science 321:836–838. https://doi.org/10.1126/science.1159023
Boualem A, Troadec C, Camps C, Lemhemdi A, Morin H, Sari MA, Fraenkel-Zagouri R, Kovalski I, Dogimont C, Perl-Treves R, Bendahmane A (2015) A cucurbit androecy gene reveals how unisexual flowers develop and dioecy emerges. Science 350:688–691. https://doi.org/10.1126/science.aac8370
Boualem A, Troadec C, Kovalski I, Sari MA, Perl-Treves R, Bendahmane A (2009) A conserved ethylene biosynthesis enzyme leads to andromonoecy in two cucumis species. PloS One 4:e6144. https://doi.org/10.1371/journal.pone.0006144
Byers RE, Baker LR, Sell HM, Herner RC, Dilley DR (1972) Ethylene: a natural regulator of sex expression of cucumis melo L. Proc Natl Acad Sci U S A 69:717–720. https://doi.org/10.1073/pnas.69.3.717
Chen C, Chen H, Zhang Y, Thomas HR, Frank MH, He Y, **a R (2020) TBtools: an integrative toolkit developed for interactive analyses of big biological data. Mol Plant 13:1194–1202. https://doi.org/10.1016/j.molp.2020.06.009
Chen H, Sun J, Li S, Cui Q, Zhang H, **n F, Wang H, Lin T, Gao D, Wang S, Li X, Wang D, Zhang Z, Xu Z, Huang S (2016) An acc oxidase gene essential for cucumber carpel development. Mol Plant 9:1315–1327. https://doi.org/10.1016/j.molp.2016.06.018
Chen Y, Chen Y, Shi C, Huang Z, Zhang Y, Li S, Li Y, Ye J, Yu C, Li Z, Zhang X, Wang J, Yang H, Fang L, Chen Q (2018) SOAPnuke: a mapreduce acceleration-supported software for integrated quality control and preprocessing of high-throughput sequencing data. Gigascience 7:1–6. https://doi.org/10.1093/gigascience/gix120
Cheng H, Song S, **ao L, Soo HM, Cheng Z, **e D, Peng J (2009) Gibberellin acts through jasmonate to control the expression of MYB21, MYB24, and MYB57 to promote stamen filament growth in arabidopsis. PLoS Genet 5:e1000440. https://doi.org/10.1371/journal.pgen.1000440
Cui J, Luo S, Niu Y, Huang R, Wen Q, Su J, Miao N, He W, Dong Z, Cheng J, Hu K (2018) A RAD-based genetic map for anchoring scaffold sequences and identifying QTLs in bitter gourd (Momordica charantia). Front Plant Sci 9:477. https://doi.org/10.3389/fpls.2018.00477
Cui J, Yang Y, Luo S, Wang L, Huang R, Wen Q, Han X, Miao N, Cheng J, Liu Z, Zhang C, Feng C, Zhu H, Su J, Wan X, Hu F, Niu Y, Zheng X, Yang Y, Shan D, Dong Z, He W, Dhillon NPS, Hu K (2020) Whole-genome sequencing provides insights into the genetic diversity and domestication of bitter gourd (Momordica spp.). Hortic Res 7:85. https://doi.org/10.1038/s41438-020-0305-5
Dai D, Wang L, Liu Y, Chu M, Wang J, Ji P, Sheng Y (2022) Screening of key genes promoting stamen formation induced by silver nitrate in gynoecious melon. https://doi.org/10.2139/ssrn.4019483
Daumann M, Hickl D, Zimmer D, DeTar RA, Kunz HH, Möhlmann T (2018) Characterization of filament-forming CTP synthases from Arabidopsis thaliana. Plant J 96:316–328. https://doi.org/10.1111/tpj.14032
Delcher AL, Salzberg SL, Phillippy AM (2003) Using MUMmer to identify similar regions in large sequence sets. Curr Protoc Bioinformatics 00:1–18. https://doi.org/10.1002/0471250953.bi1003s00
Dellaporta SL, Calderon-Urrea A (1993) Sex determination in flowering plants. Plant Cell 5:1241–1251. https://doi.org/10.1105/tpc.5.10.1241
Eishin I, Toyoaki I (2005) Bisexual flower induction by the application of silver nitrate in Gynoecious Balsam Pear (Momordica charantia L.). Horticultural Research (japan) 4:391–395. https://doi.org/10.2503/hrj.4.391
Feng B, Lu D, Ma X, Peng Y, Sun Y, Ning G, Ma H (2012) Regulation of the arabidopsis anther transcriptome by DYT1 for pollen development. Plant J 72:612–624. https://doi.org/10.1111/j.1365-313X.2012.05104.x
Flanagan CA, Hu Y, Ma H (1996) Specific expression of the AGL1 MADS-box gene suggests regulatory functions in Arabidopsis gynoecium and ovule development. Plant J 10:343–353. https://doi.org/10.1046/j.1365-313x.1996.10020343.x
Florea L, Song L, Salzberg SL (2013) Thousands of exon skip** events differentiate among splicing patterns in sixteen human tissues. F1000Res 2:188. https://doi.org/10.12688/f1000research.2-188.v2
Fujii S, Toriyama K (2005) Molecular map** of the fertility restorer gene for ms-CW-type cytoplasmic male sterility of rice. Theor Appl Genet 111:696–701. https://doi.org/10.1007/s00122-005-2054-0
Fujii S, Toriyama K (2009) Suppressed expression of retrograde-regulated male sterility restores pollen fertility in cytoplasmic male sterile rice plants. Proc Natl Acad Sci U S A 106:9513–9518. https://doi.org/10.1073/pnas.0901860106
Gangadhara Rao P, Behera TK, Gaikwad AB, Munshi AD, Jat GS, Boopalakrishnan G (2018) Map** and QTL analysis of gynoecy and earliness in bitter gourd (Momordica charantia L) Using genoty**-by-sequencing (GBS) technology. Front Plant Sci 9:1555. https://doi.org/10.3389/fpls.2018.01555
Gopalan G, He Z, Battaile KP, Luan S, Swaminathan K (2006) Structural comparison of oxidized and reduced FKBP13 from arabidopsis thaliana. Proteins 65:789–795. https://doi.org/10.1002/prot.21108
Hamrick JL, Godt MJW (1996) Effects of life history traits on genetic diversity in plant species. PHILOS T R SOC B 351:1291–1298. https://doi.org/10.1098/rstb.1996.0112
Hickl D, Scheuring D, Möhlmann T (2021) CTP synthase 2 from arabidopsis thaliana Is required for complete embryo development. Front Plant Sci 12:652434. https://doi.org/10.3389/fpls.2021.652434
Hill JT, Demarest BL, Bisgrove BW, Gorsi B, Su YC, Yost HJ (2013) MMAPPR: mutation map** analysis pipeline for pooled RNA-seq. Genome Res 23:687–697. https://doi.org/10.1101/gr.146936.112
Hu B, Li D, Liu X, Qi J, Gao D, Zhao S, Huang S, Sun J, Yang L (2017) Engineering non-transgenic gynoecious cucumber using an improved transformation protocol and optimized CRISPR/Cas9 system. Mol Plant 10:1575–1578. https://doi.org/10.1016/j.molp.2017.09.005
Ji G, Zhang J, Zhang H, Sun H, Gong G, Shi J, Tian S, Guo S, Ren Y, Shen H, Gao J, Xu Y (2016) Mutation in the gene encoding 1-aminocyclopropane-1-carboxylate synthase 4 (CitACS4) led to andromonoecy in watermelon. J Integr Plant Biol 58:762–765. https://doi.org/10.1111/jipb.12466
Kim D, Langmead B, Salzberg SL (2015) HISAT: a fast spliced aligner with low memory requirements. Nat Methods 12:357–360. https://doi.org/10.1038/nmeth.3317
Kole C (2020) The bitter gourd genome. Springer, Cham Switzerland
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 337:1870–1874. https://doi.org/10.1093/molbev/msw054
Leclercq J, Adams-Phillips LC, Zegzouti H, Jones B, Latché A, Giovannoni JJ, Pech JC, Bouzayen M (2002) LeCTR1, a tomato CTR1-like gene, demonstrates ethylene signaling ability in Arabidopsis and novel expression patterns in tomato. Plant Physiol 130:1132–1142. https://doi.org/10.1104/pp.009415
Lewis D (1942) The evolution of sex in flowering plants. Biol Rev 17:46–67. https://doi.org/10.1111/j.1469-185X.1942.tb00431.x
Li H, Durbin R (2009) Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25:1754–1760. https://doi.org/10.1093/bioinformatics/btp324
Li Z, Han Y, Niu H, Wang Y, Jiang B, Weng Y (2020) Gynoecy instability in cucumber (Cucumis sativus L.) is due to unequal crossover at the copy number variation-dependent femaleness (F) locus. Hortic Res 7:32. https://doi.org/10.1038/s41438-020-0251-2
Li Z, Huang S, Liu S, Pan J, Zhang Z, Tao Q, Shi Q, Jia Z, Zhang W, Chen H, Si L, Zhu L, Cai R (2009) Molecular isolation of the M gene suggests that a conserved-residue conversion induces the formation of bisexual flowers in cucumber plants. Genetics 182:1381–1385. https://doi.org/10.1534/genetics.109.104737
Liu JL (2010) Intracellular compartmentation of CTP synthase in Drosophila. J Genet Genomics 37:281–296. https://doi.org/10.1016/s1673-8527(09)60046-1
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 25:402–408. https://doi.org/10.1006/meth.2001.1262
Love MI, Huber W, Anders S (2014) Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 15:550. https://doi.org/10.1186/s13059-014-0550-8
Lynch EM, Hicks DR, Shepherd M, Endrizzi JA, Maker A, Hansen JM, Barry RM, Gitai Z, Baldwin EP, Kollman JM (2017) Human CTP synthase filament structure reveals the active enzyme conformation. Nat Struct Mol Biol 24:507–514. https://doi.org/10.1038/nsmb.3407
Macdonald PM, Struhl G (1988) Cis-acting sequences responsible for anterior localization of bicoid mRNA in Drosophila embryos. Nature 336:595–598. https://doi.org/10.1038/336595a0
Mandaokar A, Browse J (2009) MYB108 acts together with MYB24 to regulate jasmonate-mediated stamen maturation in Arabidopsis. Plant Physiol 149:851–862. https://doi.org/10.1104/pp.108.132597
Manzano S, Aguado E, Martínez C, Megías Z, García A, Jamilena M (2016) The Ethylene Biosynthesis Gene CitACS4 Regulates Monoecy/Andromonoecy in Watermelon (Citrullus lanatus). PloS one 11:e0154362. https://doi.org/10.1371/journal.pone.0154362
Manzano S, Martínez C, García JM, Megías Z, Jamilena M (2014) Involvement of ethylene in sex expression and female flower development in watermelon (Citrullus lanatus). Plant Physiol Biochem 85:96–104. https://doi.org/10.1016/j.plaphy.2014.11.004
Manzano S, Martínez C, Megías Z, Gómez P, Garrido D, Jamilena M (2011) The role of ethylene and brassinosteroids in the control of sex expression and flower development in Cucurbita pepo. Plant Growth Regul 65:213–221. https://doi.org/10.1007/s10725-011-9589-7
Mao X, Cai T, Olyarchuk JG, Wei L (2005) Automated genome annotation and pathway identification using the KEGG Orthology (KO) as a controlled vocabulary. Bioinformatics 21:3787–3793. https://doi.org/10.1093/bioinformatics/bti430
Martin A, Troadec C, Boualem A, Rajab M, Fernandez R, Morin H, Pitrat M, Dogimont C, Bendahmane A (2009) A transposon-induced epigenetic change leads to sex determination in melon. Nature 461:1135–1138. https://doi.org/10.1038/nature08498
Matsumura H, Hsiao MC, Lin YP, Toyoda A, Taniai N, Tarora K, Urasaki N, Anand SS, Dhillon NPS, Schafleitner R, Lee CR (2020) Long-read bitter gourd (Momordica charantia) genome and the genomic architecture of nonclassic domestication. Proc Natl Acad Sci U S A 117:14543–14551. https://doi.org/10.1073/pnas.1921016117
Matsumura H, Miyagi N, Taniai N, Fukushima M, Tarora K, Shudo A, Urasaki N (2014) Map** of the gynoecy in bitter gourd (Momordica charantia) using RAD-seq analysis. PloS one 9:e87138. https://doi.org/10.1371/journal.pone.0087138
Mayr C (2019) What Are 3′ UTRs Doing? Cold Spring Harb Perspect Biol 11:a034728. https://doi.org/10.1101/cshperspect.a034728
McKenna A, Hanna M, Banks E, Sivachenko A, Cibulskis K, Kernytsky A, Garimella K, Altshuler D, Gabriel S, Daly M, DePristo MA (2010) The genome analysis toolkit: a mapreduce framework for analyzing next-generation DNA sequencing data. Genome Res 20:1297–1303. https://doi.org/10.1101/gr.107524.110
Mibus H, Tatlioglu T (2004) Molecular characterization and isolation of the F/f gene for femaleness in cucumber (Cucumis sativus L.). Theor Appl Genet 109:1669–1676. https://doi.org/10.1007/s00122-004-1793-7
Noree C, Monfort E, Shiau AK, Wilhelm JE (2014) Common regulatory control of CTP synthase enzyme activity and filament formation. Mol Biol Cell 25:2282–2290. https://doi.org/10.1091/mbc.E14-04-0912
Pertea M, Pertea GM, Antonescu CM, Chang TC, Mendell JT, Salzberg SL (2015) StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat Biotechnol 33:290–295. https://doi.org/10.1038/nbt.3122
Porebski S, Bailey LG, Baum BR (1997) Modification of a CTAB DNA extraction protocol for plants containing high polysaccharide and polyphenol components. Plant Mol Biol Rep 15:8–15. https://doi.org/10.1007/BF02772108
Ram D, Kumar S, Singh M, Rai M, Kalloo G (2006) Inheritance of gynoecism in bitter gourd (Momordica charantia L.). J Hered 97:294–295. https://doi.org/10.1093/jhered/esj028
Rudich J, Halevy AH, Kedar N (1969) Increase in femaleness of three cucurbits by treatment with ethrel, an ethylene releasing compound. Planta 86:69–76. https://doi.org/10.1007/bf00385305
Schaefer H, Renner SS (2010) A three-genome phylogeny of Momordica (Cucurbitaceae) suggests seven returns from dioecy to monoecy and recent long-distance dispersal to Asia. Mol Phylogenet Evol 54:553–560. https://doi.org/10.1016/j.ympev.2009.08.006
Schaefer H, Renner SS (2011) Cucurbitaceae. In: Kubitzki K (ed) Flowering plants eudicots: sapindales, cucurbitales, myrtaceae. Springer, Berlin Heidelberg, pp 112–174. https://doi.org/10.1007/978-3-642-14397-7_10
Siepel A, Bejerano G, Pedersen JS, Hinrichs AS, Hou M, Rosenbloom K, Clawson H, Spieth J, Hillier LW, Richards S, Weinstock GM, Wilson RK, Gibbs RA, Kent WJ, Miller W, Haussler D (2005) Evolutionarily conserved elements in vertebrate, insect, worm, and yeast genomes. Genome Res 15:1034–1050. https://doi.org/10.1101/gr.3715005
Tan J, Tao Q, Niu H, Zhang Z, Li D, Gong Z, Weng Y, Li Z (2015) A novel allele of monoecious (m) locus is responsible for elongated fruit shape and perfect flowers in cucumber (Cucumis sativus L.). Theor Appl Genet 128:2483–2493. https://doi.org/10.1007/s00122-015-2603-0
Tao Q, Niu H, Wang Z, Zhang W, Wang H, Wang S, Zhang X, Li Z (2018) Ethylene responsive factor ERF110 mediates ethylene-regulated transcription of a sex determination-related orthologous gene in two Cucumis species. J Exp Bot 69:2953–2965. https://doi.org/10.1093/jxb/ery128
Van Ooijen JW (2006) Joinmap4, software for calculation of genetic linkage maps in experimental populations. Wageningen, Kyazma BV
Wang K, Li M, Hakonarson H (2010) ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res 38:e164. https://doi.org/10.1093/nar/gkq603
Wang L, Ko EE, Tran J, Qiao H (2020) TREE1-EIN3-mediated transcriptional repression inhibits shoot growth in response to ethylene. Proc Natl Acad Sci U S A 117:29178–29189. https://doi.org/10.1073/pnas.2018735117
Wang S, Li Z, Yang G, Ho CT, Li S (2017) Momordica charantia: a popular health-promoting vegetable with multifunctionality. Food Funct 8:1749–1762. https://doi.org/10.1039/c6fo01812b
**e X, Lu J, Kulbokas EJ, Golub TR, Mootha V, Lindblad-Toh K, Lander ES, Kellis M (2005) Systematic discovery of regulatory motifs in human promoters and 3’ UTRs by comparison of several mammals. Nature 434:338–345. https://doi.org/10.1038/nature03441
Yamasaki S, Manabe K (2011) Application of silver nitrate induces functional bisexual flowers in gynoecious cucumber plants (Cucumis sativus L.). J JPN SOC HORTIC SCI 80:66–75. https://doi.org/10.2503/jjshs1.80.66
Yoon J, Cho LH, Kim SR, Tun W, Peng X, Pasriga R, Moon S, Hong WJ, Ji H, Jung KH, Jeon JS, An G (2021) CTP synthase is essential for early endosperm development by regulating nuclei spacing. Plant Biotechnol J 19:2177–2191. https://doi.org/10.1111/pbi.13644
Zhang J, Guo S, Ji G, Zhao H, Sun H, Ren Y, Tian S, Li M, Gong G, Zhang H, Xu Y (2020a) A unique chromosome translocation disrupting ClWIP1 leads to gynoecy in watermelon. Plant J 101:265–277. https://doi.org/10.1111/tpj.14537
Zhang J, Shi J, Ji G, Zhang H, Gong G, Guo S, Ren Y, Fan J, Tian S, Xu Y (2017) Modulation of sex expression in four forms of watermelon by gibberellin, ethephone and silver nitrate. Hortic Plant J 3:91–100. https://doi.org/10.1016/j.hpj.2017.07.010
Zhang YC, He RR, Lian JP, Zhou YF, Zhang F, Li QF, Yu Y, Feng YZ, Yang YW, Lei MQ, He H, Zhang Z, Chen YQ (2020b) OsmiR528 regulates rice-pollen intine formation by targeting an uclacyanin to influence flavonoid metabolism. Proc Natl Acad Sci U S A 117:727–732. https://doi.org/10.1073/pnas.1810968117
Funding
This work was supported by the Key Project of Basic and Applied Research for University in Guangdong Province (2018KZDXM016), the Science and Technology Planning Project of Guangdong Province (2018B020202007), the Guangzhou Science and Technology Plan Projects (202002020086, 202102020800, and 202206010170), and the Science and Technology Plan Projects of Guangdong Province (2019A050520002).
Author information
Authors and Affiliations
Contributions
KH and JWC conceived and designed the research. JZ and JJC performed most of the experiments and wrote the manuscript. JL and CFZ performed statistical analysis. FH and JCD provided helpful discussions. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Ethical approval
The authors declare that this study complies with the current laws of China.
Additional information
Communicated by Yiqun Weng.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhong, J., Cui, J., Liu, J. et al. Fine-map** and candidate gene analysis of the Mcgy1 locus responsible for gynoecy in bitter gourd (Momordica spp.). Theor Appl Genet 136, 81 (2023). https://doi.org/10.1007/s00122-023-04314-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00122-023-04314-0