Abstract
Oxysterol-binding protein/OSBP-related proteins (ORPs) constitute a conserved family of sterol/phospholipid-binding proteins with lipid transporter or sensor functions. We investigated the spatial occurrence and regulation of the interactions of human OSBP/ORPs or the S. cerevisiae orthologs, the Osh (OSBP homolog) proteins, with their endoplasmic reticulum (ER) anchors, the VAMP-associated proteins (VAPs), by employing bimolecular fluorescence complementation and pull-down set-ups. The ORP–VAP interactions localize frequently at distinct subcellular sites, shown in several cases to represent membrane contact sites (MCSs). Using established ORP ligand-binding domain mutants and pull-down assays with recombinant proteins, we show that ORP liganding regulates the ORP–VAP association, alters the subcellular targeting of ORP–VAP complexes, or modifies organelle morphology. There is distinct protein specificity in the effects of the mutants on subcellular targeting of ORP–VAP complexes. We provide evidence that complexes of human ORP2 and VAPs at ER–lipid droplet interfaces regulate the hydrolysis of triglycerides and lipid droplet turnover. The data suggest evolutionarily conserved, complex ligand-dependent functions of ORP–VAP complexes at MCSs, with implications for cellular lipid homeostasis and signaling.








Similar content being viewed by others
Abbreviations
- BiFC:
-
Bimolecular fluorescence complementation
- EM:
-
Electron microscopy
- ER:
-
Endoplasmic reticulum
- FFAT:
-
Two phenylalanines in an acidic tract
- GST:
-
Glutathione-S-transferase
- LD:
-
Lipid droplet
- MCS:
-
Membrane contact site
- OHC:
-
Hydroxycholesterol
- ORD:
-
OSBP-related ligand-binding domain
- ORP:
-
OSBP-related protein
- OSBP:
-
Oxysterol-binding protein
- Osh:
-
OSBP homolog (in yeast)
- PI4P:
-
Phosphatidylinositol-4-phosphate
- PIP:
-
Phosphoinositide
- PM:
-
Plasma membrane
- TG:
-
Triglyceride
- VAMP:
-
Vesicle-associated membrane protein
- VAP:
-
VAMP-associated protein
References
Dawson PA, Ridgway ND, Slaughter CA, Brown MS, Goldstein JL (1989) cDNA cloning and expression of oxysterol-binding protein, an oligomer with a potential leucine zipper. J Biol Chem 264:16798–16803
Taylor FR, Saucier SE, Shown EP, Parish EJ, Kandutsch AA (1984) Correlation between oxysterol binding to a cytosolic binding protein and potency in the repression of hydroxymethylglutaryl coenzyme A reductase. J Biol Chem 259:12382–12387
Wang PY, Weng J, Anderson RG (2005) OSBP is a cholesterol-regulated scaffolding protein in control of ERK 1/2 activation. Science 307:1472–1476
Mesmin B, Bigay J, Moser von Filseck J, Lacas-Gervais S, Drin G, Antonny B (2013) A four-step cycle driven by PI(4)P hydrolysis directs sterol/PI(4)P exchange by the ER-Golgi tether OSBP. Cell 155:830–843
Anniss AM, Apostolopoulos J, Dworkin S, Purton LE, Sparrow RL (2002) An oxysterol-binding protein family identified in the mouse. DNA Cell Biol 21:571–580
Jaworski CJ, Moreira E, Li A, Lee R, Rodriguez IR (2001) A family of 12 human genes containing oxysterol-binding domains. Genomics 78:185–196
Lehto M, Laitinen S, Chinetti G, Johansson M, Ehnholm C, Staels B, Ikonen E, Olkkonen VM (2001) The OSBP-related protein family in humans. J Lipid Res 42:1203–1213
Beh CT, Cool L, Phillips J, Rine J (2001) Overlap** functions of the yeast oxysterol-binding protein homologues. Genetics 157:1117–1140
Olkkonen VM, Li S (2013) Oxysterol-binding proteins: sterol and phosphoinositide sensors coordinating transport, signaling and metabolism. Prog Lipid Res 52:529–538
Raychaudhuri S, Prinz WA (2010) The diverse functions of oxysterol-binding proteins. Annu Rev Cell Dev Biol 26:157–177
Ridgway ND (2010) Oxysterol-binding proteins. Subcell Biochem 51:159–182
Kaiser SE, Brickner JH, Reilein AR, Fenn TD, Walter P, Brunger AT (2005) Structural basis of FFAT motif-mediated ER targeting. Structure 13:1035–1045
Loewen CJ, Roy A, Levine TP (2003) A conserved ER targeting motif in three families of lipid binding proteins and in Opi1p binds VAP. EMBO J 22:2025–2035
Du X, Kumar J, Ferguson C, Schulz TA, Ong YS, Hong W, Prinz WA, Parton RG, Brown AJ, Yang H (2011) A role for oxysterol-binding protein-related protein 5 in endosomal cholesterol trafficking. J Cell Biol 192:121–135
Yan D, Mäyränpää MI, Wong J, Perttilä J, Lehto M, Jauhiainen M, Kovanen PT, Ehnholm C, Brown AJ, Olkkonen VM (2008) OSBP-related protein 8 (ORP8) suppresses ABCA1 expression and cholesterol efflux from macrophages. J Biol Chem 283:332–340
Kagiwada S, Hosaka K, Murata M, Nikawa J, Takatsuki A (1998) The Saccharomyces cerevisiae SCS2 gene product, a homolog of a synaptobrevin-associated protein, is an integral membrane protein of the endoplasmic reticulum and is required for inositol metabolism. J Bacteriol 180:1700–1708
Nishimura Y, Hayashi M, Inada H, Tanaka T (1999) Molecular cloning and characterization of mammalian homologues of vesicle-associated membrane protein-associated (VAMP-associated) proteins. Biochem Biophys Res Commun 254:21–26
Skehel PA, Armitage BA, Bartsch D, Hu Y, Kaang BK, Siegelbaum SA, Kandel ER, Martin KC (1995) Proteins functioning in synaptic transmission at the sensory to motor synapse of Aplysia. Neuropharmacology 34:1379–1385
Skehel PA, Fabian-Fine R, Kandel ER (2000) Mouse VAP33 is associated with the endoplasmic reticulum and microtubules. Proc Natl Acad Sci USA 97:1101–1106
Lev S, Ben Halevy D, Peretti D, Dahan N (2008) The VAP protein family: from cellular functions to motor neuron disease. Trends Cell Biol 18:282–290
Moustaqim-Barrette A, Lin YQ, Pradhan S, Neely GG, Bellen HJ, Tsuda H (2014) The amyotrophic lateral sclerosis 8 protein, VAP, is required for ER protein quality control. Hum Mol Genet 23:1975–1989
Nishimura AL, Al-Chalabi A, Zatz M (2005) A common founder for amyotrophic lateral sclerosis type 8 (ALS8) in the Brazilian population. Hum Genet 118:499–500
Nishimura AL, Mitne-Neto M, Silva HC, Richieri-Costa A, Middleton S, Cascio D, Kok F, Oliveira JR, Gillingwater T, Webb J, Skehel P, Zatz M (2004) A mutation in the vesicle-trafficking protein VAPB causes late-onset spinal muscular atrophy and amyotrophic lateral sclerosis. Am J Hum Genet 75:822–831
Kvam E, Goldfarb DS (2004) Nvj1p is the outer-nuclear-membrane receptor for oxysterol-binding protein homolog Osh1p in Saccharomyces cerevisiae. J Cell Sci 117:4959–4968
Levine TP, Munro S (2001) Dual targeting of Osh1p, a yeast homologue of oxysterol-binding protein, to both the Golgi and the nucleus–vacuole junction. Mol Biol Cell 12:1633–1644
Rocha N, Kuijl C, van der Kant R, Janssen L, Houben D, Janssen H, Zwart W, Neefjes J (2009) Cholesterol sensor ORP1L contacts the ER protein VAP to control Rab7-RILP-p150 Glued and late endosome positioning. J Cell Biol 185:1209–1225
Schulz TA, Choi MG, Raychaudhuri S, Mears JA, Ghirlando R, Hinshaw JE, Prinz WA (2009) Lipid-regulated sterol transfer between closely apposed membranes by oxysterol-binding protein homologues. J Cell Biol 187:889–903
Stefan CJ, Manford AG, Emr SD (2013) ER-PM connections: sites of information transfer and inter-organelle communication. Curr Opin Cell Biol 25:434–442
Tavassoli S, Chao JT, Young BP, Cox RC, Prinz WA, de Kroon AI, Loewen CJ (2013) Plasma membrane–endoplasmic reticulum contact sites regulate phosphatidylcholine synthesis. EMBO Rep 14:434–440
Vihervaara T, Uronen RL, Wohlfahrt G, Björkhem I, Ikonen E, Olkkonen VM (2011) Sterol binding by OSBP-related protein 1L regulates late endosome motility and function. Cell Mol Life Sci 68:537–551
Stefan CJ, Manford AG, Baird D, Yamada-Hanff J, Mao Y, Emr SD (2011) Osh proteins regulate phosphoinositide metabolism at ER-plasma membrane contact sites. Cell 144:389–401
Elbaz Y, Schuldiner M (2011) Staying in touch: the molecular era of organelle contact sites. Trends Biochem Sci 36:616–623
Helle SC, Kanfer G, Kolar K, Lang A, Michel AH, Kornmann B (2013) Organization and function of membrane contact sites. Biochim Biophys Acta 1833:2526–2541
Klecker T, Bockler S, Westermann B (2014) Making connections: interorganelle contacts orchestrate mitochondrial behavior. Trends Cell Biol (Epub ahead of print)
Levine T, Loewen C (2006) Inter-organelle membrane contact sites: through a glass, darkly. Curr Opin Cell Biol 18:371–378
Toulmay A, Prinz WA (2011) Lipid transfer and signaling at organelle contact sites: the tip of the iceberg. Curr Opin Cell Biol 23:458–463
Blanchette-Mackie EJ, Dwyer NK, Barber T, Coxey RA, Takeda T, Rondinone CM, Theodorakis JL, Greenberg AS, Londos C (1995) Perilipin is located on the surface layer of intracellular lipid droplets in adipocytes. J Lipid Res 36:1211–1226
Robenek H, Hofnagel O, Buers I, Robenek MJ, Troyer D, Severs NJ (2006) Adipophilin-enriched domains in the ER membrane are sites of lipid droplet biogenesis. J Cell Sci 119:4215–4224
Xu N, Zhang SO, Cole RA, McKinney SA, Guo F, Haas JT, Bobba S, Farese RV Jr, Mak HY (2012) The FATP1–DGAT2 complex facilitates lipid droplet expansion at the ER-lipid droplet interface. J Cell Biol 198:895–911
Soni KG, Mardones GA, Sougrat R, Smirnova E, Jackson CL, Bonifacino JS (2009) Coatomer-dependent protein delivery to lipid droplets. J Cell Sci 122:1834–1841
Wilfling F, Thiam AR, Olarte MJ, Wang J, Beck R, Gould TJ, Allgeyer ES, Pincet F, Bewersdorf J, Farese RV Jr, Walther TC (2014) Arf1/COPI machinery acts directly on lipid droplets and enables their connection to the ER for protein targeting. Elife 3:e01607
Wilfling F, Wang H, Haas JT, Krahmer N, Gould TJ, Uchida A, Cheng JX, Graham M, Christiano R, Frohlich F, Liu X, Buhman KK, Coleman RA, Bewersdorf J, Farese RV Jr, Walther TC (2013) Triacylglycerol synthesis enzymes mediate lipid droplet growth by relocalizing from the ER to lipid droplets. Dev Cell 24:384–399
de Brito OM, Scorrano L (2008) Mitofusin 2 tethers endoplasmic reticulum to mitochondria. Nature 456:605–610
Giordano F, Saheki Y, Idevall-Hagren O, Colombo SF, Pirruccello M, Milosevic I, Gracheva EO, Bagriantsev SN, Borgese N, De Camilli P (2013) PI(4,5)P(2)-dependent and Ca(2+)-regulated ER-PM interactions mediated by the extended synaptotagmins. Cell 153:1494–1509
Kornmann B, Currie E, Collins SR, Schuldiner M, Nunnari J, Weissman JS, Walter P (2009) An ER–mitochondria tethering complex revealed by a synthetic biology screen. Science 325:477–481
Manford AG, Stefan CJ, Yuan HL, Macgurn JA, Emr SD (2012) ER-to-plasma membrane tethering proteins regulate cell signaling and ER morphology. Dev Cell 23:1129–1140
Srikanth S, Jew M, Kim KD, Yee MK, Abramson J, Gwack Y (2012) Junctate is a Ca2+-sensing structural component of Orai1 and stromal interaction molecule 1 (STIM1). Proc Natl Acad Sci USA 109:8682–8687
Takeshima H, Komazaki S, Nishi M, Iino M, Kangawa K (2000) Junctophilins: a novel family of junctional membrane complex proteins. Mol Cell 6:11–22
Wang Y, Deng X, Mancarella S, Hendron E, Eguchi S, Soboloff J, Tang XD, Gill DL (2010) The calcium store sensor, STIM1, reciprocally controls Orai and CaV1.2 channels. Science 330:105–109
Yuan JP, Kiselyov K, Shin DM, Chen J, Shcheynikov N, Kang SH, Dehoff MH, Schwarz MK, Seeburg PH, Muallem S, Worley PF (2003) Homer binds TRPC family channels and is required for gating of TRPC1 by IP3 receptors. Cell 114:777–789
Zhou Y, Li S, Mäyränpää MI, Zhong W, Back N, Yan D, Olkkonen VM (2010) OSBP-related protein 11 (ORP11) dimerizes with ORP9 and localizes at the Golgi-late endosome interface. Exp Cell Res 316:3304–3316
Skarp KP, Zhao X, Weber M, Jäntti J (2008) Use of bimolecular fluorescence complementation in yeast Saccharomyces cerevisiae. Methods Mol Biol 457:165–175
de Saint-Jean M, Delfosse V, Douguet D, Chicanne G, Payrastre B, Bourguet W, Antonny B, Drin G (2011) Osh4p exchanges sterols for phosphatidylinositol 4-phosphate between lipid bilayers. J Cell Biol 195:965–978
Laitinen S, Lehto M, Lehtonen S, Hyvärinen K, Heino S, Lehtonen E, Ehnholm C, Ikonen E, Olkkonen VM (2002) ORP2, a homolog of oxysterol binding protein, regulates cellular cholesterol metabolism. J Lipid Res 43:245–255
Lehto M, Hynynen R, Karjalainen K, Kuismanen E, Hyvärinen K, Olkkonen VM (2005) Targeting of OSBP-related protein 3 (ORP3) to endoplasmic reticulum and plasma membrane is controlled by multiple determinants. Exp Cell Res 310:445–462
Seemann J, Jokitalo EJ, Warren G (2000) The role of the tethering proteins p115 and GM130 in transport through the Golgi apparatus in vivo. Mol Biol Cell 11:635–645
Hynynen R, Suchanek M, Spandl J, Back N, Thiele C, Olkkonen VM (2009) OSBP-related protein 2 is a sterol receptor on lipid droplets that regulates the metabolism of neutral lipids. J Lipid Res 50:1305–1315
Bligh EG, Dyer WJ (1959) A rapid method of total lipid extraction and purification. Can J Biochem Physiol 37:911–917
Sherman F (1991) Getting started with yeast. Methods Enzymol 194:3–21
Novick P, Field C, Schekman R (1980) Identification of 23 complementation groups required for post-translational events in the yeast secretory pathway. Cell 21:205–215
Kerppola TK (2008) Bimolecular fluorescence complementation (BiFC) analysis as a probe of protein interactions in living cells. Annu Rev Biophys 37:465–487
Ngo M, Ridgway ND (2009) Oxysterol binding protein-related Protein 9 (ORP9) is a cholesterol transfer protein that regulates Golgi structure and function. Mol Biol Cell 20:1388–1399
Suchanek M, Hynynen R, Wohlfahrt G, Lehto M, Johansson M, Saarinen H, Radzikowska A, Thiele C, Olkkonen VM (2007) The mammalian OSBP-related proteins (ORP) bind 25-hydroxycholesterol in an evolutionarily conserved pocket. Biochem J 405:473–480
Wyles JP, Perry RJ, Ridgway ND (2007) Characterization of the sterol-binding domain of oxysterol-binding protein (OSBP)-related protein 4 reveals a novel role in vimentin organization. Exp Cell Res 313:1426–1437
Perry RJ, Ridgway ND (2006) Oxysterol-binding protein and vesicle-associated membrane protein-associated protein are required for sterol-dependent activation of the ceramide transport protein. Mol Biol Cell 17:2604–2616
Nhek S, Ngo M, Yang X, Ng MM, Field SJ, Asara JM, Ridgway ND, Toker A (2010) Regulation of oxysterol-binding protein Golgi localization through protein kinase D-mediated phosphorylation. Mol Biol Cell 21:2327–2337
Ridgway ND, Lagace TA, Cook HW, Byers DM (1998) Differential effects of sphingomyelin hydrolysis and cholesterol transport on oxysterol-binding protein phosphorylation and Golgi localization. J Biol Chem 273:31621–31628
Tong J, Yang H, Yang H, Eom SH, Im YJ (2013) Structure of Osh3 reveals a conserved mode of phosphoinositide binding in oxysterol-binding proteins. Structure 21:1203–1213
Maeda K, Anand K, Chiapparino A, Kumar A, Poletto M, Kaksonen M, Gavin AC (2013) Interactome map uncovers phosphatidylserine transport by oxysterol-binding proteins. Nature 501:257–261
Zhou Y, Wohlfahrt G, Paavola J, Olkkonen VM (2014) A vertebrate model for the study of lipid binding/transfer protein function: conservation of OSBP-related proteins between zebrafish and human. Biochem Biophys Res Commun 446:675–680
Liu X, Ridgway ND (2014) Characterization of the sterol and phosphatidylinositol 4-phosphate binding properties of Golgi-associated OSBP-related protein 9 (ORP9). PLoS ONE 9:e108368
Nissilä E, Ohsaki Y, Weber-Boyvat M, Perttilä J, Ikonen E, Olkkonen VM (2012) ORP10, a cholesterol binding protein associated with microtubules, regulates apolipoprotein B-100 secretion. Biochim Biophys Acta 1821:1472–1484
Perttilä J, Merikanto K, Naukkarinen J, Surakka I, Martin NW, Tanhuanpää K, Grimard V, Taskinen MR, Thiele C, Salomaa V, Jula A, Perola M, Virtanen I, Peltonen L, Olkkonen VM (2009) OSBPL10, a novel candidate gene for high triglyceride trait in dyslipidemic Finnish subjects, regulates cellular lipid metabolism. J Mol Med 87:825–835
Pennetta G, Hiesinger PR, Fabian-Fine R, Meinertzhagen IA, Bellen HJ (2002) Drosophila VAP-33A directs bouton formation at neuromuscular junctions in a dosage-dependent manner. Neuron 35:291–306
Johansson M, Bocher V, Lehto M, Chinetti G, Kuismanen E, Ehnholm C, Staels B, Olkkonen VM (2003) The two variants of oxysterol binding protein-related protein-1 display different tissue expression patterns, have different intracellular localization, and are functionally distinct. Mol Biol Cell 14:903–915
Lagace TA, Byers DM, Cook HW, Ridgway ND (1997) Altered regulation of cholesterol and cholesteryl ester synthesis in Chinese-hamster ovary cells overexpressing the oxysterol-binding protein is dependent on the pleckstrin homology domain. Biochem J 326:205–213
Wyles JP, Ridgway ND (2004) VAMP-associated protein-A regulates partitioning of oxysterol-binding protein-related protein-9 between the endoplasmic reticulum and Golgi apparatus. Exp Cell Res 297:533–547
Olkkonen VM, Levine TP (2004) Oxysterol binding proteins: in more than one place at one time? Biochem Cell Biol 82:87–98
Lagace TA, Byers DM, Cook HW, Ridgway ND (1999) Chinese hamster ovary cells overexpressing the oxysterol binding protein (OSBP) display enhanced synthesis of sphingomyelin in response to 25-hydroxycholesterol. J Lipid Res 40:109–116
Peretti D, Dahan N, Shimoni E, Hirschberg K, Lev S (2008) Coordinated lipid transfer between the endoplasmic reticulum and the Golgi complex requires the VAP proteins and is essential for Golgi-mediated transport. Mol Biol Cell 19:3871–3884
Burgett AW, Poulsen TB, Wangkanont K, Anderson DR, Kikuchi C, Shimada K, Okubo S, Fortner KC, Mimaki Y, Kuroda M, Murphy JP, Schwalb DJ, Petrella EC, Cornella-Taracido I, Schirle M, Tallarico JA, Shair MD (2011) Natural products reveal cancer cell dependence on oxysterol-binding proteins. Nat Chem Biol 7:639–647
Charman M, Colbourne TR, Pietrangelo A, Kreplak L, Ridgway ND (2014) Oxysterol-binding protein (OSBP)-related protein 4 (ORP4) is essential for cell proliferation and survival. J Biol Chem 289:15705–15717
Bonifacino JS, Glick BS (2004) The mechanisms of vesicle budding and fusion. Cell 116:153–166
Alfaro G, Johansen J, Dighe SA, Duamel G, Kozminski KG, Beh CT (2011) The sterol-binding protein Kes1/Osh4p is a regulator of polarized exocytosis. Traffic 12:1521–1536
Loewen CJ, Levine TP (2005) A highly conserved binding site in vesicle-associated membrane protein-associated protein (VAP) for the FFAT motif of lipid-binding proteins. J Biol Chem 280:14097–14104
Acknowledgments
Liisa Arala, Eeva Jääskeläinen and Riikka Kosonen are thanked for skillful technical assistance, Prof. M. A. De Matteis (Telethon Institute of Genetics and Medicine, Naples, Italy) for the rabbit VAPA and -B antibodies, and Dr. I. Kaverina (Vanderbilt Univ., Nashville, TN, USA) for the mCherry-GalT1(1–81) construct. We thank Eija Jokitalo and the Electron Microscopy Unit (Institute of Biotechnology, Helsinki, Finland) for assistance with the transmission electron microscopy. The study was supported by the Academy of Finland (Grant 257409 to M.W.-B.), the Finnish Concordia Fund (H. K.), the Sigrid Juselius Foundation, the Novo Nordisk Foundation, the Liv ovh Hälsa Foundation, the Finnish Foundation for Cardiovascular Research, and the Magnus Ehrnrooth Foundation (V. M. O.). The authors declare no conflicts of interest.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
18_2014_1786_MOESM1_ESM.tif
Supplementary Fig. S1 FFAT motif containing ORPs interact with VAPB at specific sites in the cell. Fluorescence imaging of HuH7 cells expressing different VnORP constructs in combination with VcVAPB and mCherry-VAPB. The BiFC signals indicating physical interaction/proximity of the ORPs and VAPB are shown in the top row, and merge of the BiFC and mCherry-VAPB fluorescence in the bottom row. (TIFF 10164 kb)
18_2014_1786_MOESM2_ESM.tif
Supplementary Fig. S2 Impact of inositol-phosphate-binding cleft mutations (mPIP) on the ORP–VAPA interaction. (A) HuH7 cells expressing the indicated wt or mPIP VnORP2 and VnORP4L constructs were lysed and subjected to GST-VAPA pull-down. Cell lysates and pull-downs were analyzed by Western blotting with anti-GFP antibodies. The GST-VAPA band was visualized as an unspecific band cross-reacting with the secondary antibody. (B) Quantification of three repeats of the experiment in (A). Shown are averages and standard deviations; *p < 0.05. (C and D) Fluorescence imaging of HuH7 cells expressing wt or mutant VnORP constructs [ORP4L (C), ORP2 (D)] in combination with VcVAPA and mCherry-VAPA. BiFC signal intensities were analyzed in a minimum of 100 cells per condition and normalized to the cotransfection marker mCherry-VAPA. Shown are BiFC signal intensity averages and standard deviations. (TIFF 43630 kb)
18_2014_1786_MOESM3_ESM.tif
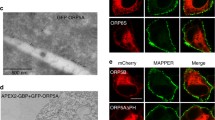
Supplementary Fig. S3 Electron microscopic analysis of cells expressing the VnORP9L and VcVAPA. (A) Transmission electron microscopy images of HuH7 cells expressing either the empty vectors (control) or VnORP9 in combination with VcVAPA. (B) Quantification of the thickness of ER sheets/tubules in control and ORP9L-VAPA-expressing cells. Shown are averages and SEM of 20 ER samples analyzed per condition; ***p < 0.005. (TIFF 43631 kb)
18_2014_1786_MOESM4_ESM.tif
Supplementary Fig. S4 Impact of the ORP2 inositol-phosphate-binding cleft mutant (mPIP) on the distribution and total area of lipid droplets (LD). HuH7 cells expressing the indicated wt or mPIP VnORP2 and VcVAPA (BiFC), with the LD (Bodipy-C12) and nuclei (DAPI) co-stained. (A) Fluorescence microscopy images displaying the three channels and their merge. (B) Quantification of total LD area in a minimum of 20 cells per condition. Shown are averages and standard deviations. (C). The LD distribution in ORP2(wt)– and ORP2(mPIP)–VAPA-expressing cells. LD distribution was analyzed in a minimum of 10 cells per condition. The center of the nucleus was used to define the cell center. Shown are averages and SEM. (TIFF 43632 kb)
Rights and permissions
About this article
Cite this article
Weber-Boyvat, M., Kentala, H., Peränen, J. et al. Ligand-dependent localization and function of ORP–VAP complexes at membrane contact sites. Cell. Mol. Life Sci. 72, 1967–1987 (2015). https://doi.org/10.1007/s00018-014-1786-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00018-014-1786-x