Abstract
Background
Previous studies have observed elevated myeloid cells in the peripheral blood of patients with Parkinson's disease (PD), but the causal relationship between them remains to be elucidated. We investigated whether there is a causal relationship between different subtypes of peripheral blood myeloid cells and PD using Mendelian randomization (MR) combined with bioinformatics analysis. Exploring the etiology of PD from the perspective of genetics can remove confounding factors and provide a more reliable theoretical basis for elucidating the pathogenesis of PD.
Methods
Comprehensive two-sample MR analysis and sensitivity analyses were conducted to explore the causal associations between 64 myeloid cell signatures and PD risk. The Venn diagram and protein-protein interaction network analysis of instrumental variables (IV) corresponding genes were used to further investigate the potential mechanism of myeloid cells influencing the pathogenesis of PD.
Results
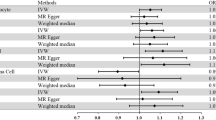
We investigated the impact of four immunophenotypes on the risk of PD, including Im MDSC% CD33dim HLA DR− CD66b− (relative count), CD33dim HLA DR+ CD11b+% CD33dim HLA DR+ (relative count), and CD11b on Mo MDSC (MFI) and CD11b on CD33br HLA DR+ CD14dim (MFI), while an immunophenotype's protective effect on PD was observed CD45 on Im MDSC (MFI). The results of bioinformatics analysis showed that CD33, NTRK2, PLD2, GRIK2 and RELN had protein interactions with the risk genes of PD.
Conclusions
Our study has demonstrated a close genetic correlation between different subtypes of myeloid cells and PD, providing guidance for early identification and immunotherapeutic development in patients with PD.





Similar content being viewed by others
Data availability
The data that supports the findings of this study are available from the authors upon reasonable request.
References
Massaquoi MS, Liguore WA, Churchill MJ, et al. Gait deficits and loss of striatal tyrosine hydroxlase/Trk-B are restored following 7,8-dihydroxyflavone treatment in a progressive MPTP mouse model of Parkinson’s Disease. Neuroscience. 2020;433:53–71.
Fasano A, Visanji NP, Liu LW, et al. Gastrointestinal dysfunction in Parkinson’s disease. Lancet Neurol. 2015;14(6):625–39.
Nair AT, Ramachandran V, Joghee NM, et al. Gut microbiota dysfunction as reliable non-invasive early diagnostic biomarkers in the pathophysiology of Parkinson’s Disease: a critical review. J Neurogastroenterol Motil. 2018;24(1):30–42.
Jankovic J, Tan EK. Parkinson’s disease: etiopathogenesis and treatment. J Neurol Neurosurg Psychiatry. 2020;91(8):795–808.
Fasciani I, Petragnano F, Aloisi G, et al. A new threat to dopamine neurons: the downside of artificial light. Neuroscience. 2020;432:216–28.
Dantzer R, O’Connor JC, Freund GG, et al. From inflammation to sickness and depression: when the immune system subjugates the brain. Nat Rev Neurosci. 2008;9(1):46–56.
Tansey MG, Wallings RL, Houser MC, et al. Inflammation and immune dysfunction in Parkinson disease. Nat Rev Immunol. 2022;22(11):657–73.
Lutters B, Foley P, Koehler PJ. The centennial lesson of encephalitis lethargica. Neurology. 2018;90(12):563–7.
Bo RX, Li YY, Zhou TT, et al. The neuroinflammatory role of glucocerebrosidase in Parkinson’s disease. Neuropharmacology. 2022;207: 108964.
Yang L, Mao K, Yu H, et al. Neuroinflammatory responses and Parkinson’ Disease: pathogenic mechanisms and therapeutic targets. J Neuroimmune Pharmacol. 2020;15(4):830–7.
Schonhoff AM, Figge DA, Williams GP, et al. Border-associated macrophages mediate the neuroinflammatory response in an alpha-synuclein model of Parkinson disease. Nat Commun. 2023;14(1):3754.
Araújo B, Caridade-Silva R, Soares-Guedes C, et al. Neuroinflammation and Parkinson’s Disease-from neurodegeneration to therapeutic opportunities. Cells. 2022;11(18):2908.
Ferrari CC, Pott Godoy MC, Tarelli R, et al. Progressive neurodegeneration and motor disabilities induced by chronic expression of IL-1beta in the substantia nigra. Neurobiol Dis. 2006;24(1):183–93.
Hagar JA, Powell DA, Aachoui Y, et al. Cytoplasmic LPS activates caspase-11: implications in TLR4-independent endotoxic shock. Science. 2013;341(6151):1250–3.
Lee E, Hwang I, Park S, et al. MPTP-driven NLRP3 inflammasome activation in microglia plays a central role in dopaminergic neurodegeneration. Cell Death Differ. 2019;26(2):213–28.
Zhou X, Lu J, Wei K, et al. Neuroprotective effect of ceftriaxone on MPTP-induced Parkinson’s disease mouse model by regulating inflammation and intestinal microbiota. Oxid Med Cell Longev. 2021;2021:9424582.
Chen S, Liu Y, Niu Y, et al. Increased abundance of myeloid-derived suppressor cells and Th17 cells in peripheral blood of newly-diagnosed Parkinson’s disease patients. Neurosci Lett. 2017;648:21–5.
Grozdanov V, Bliederhaeuser C, Ruf WP, et al. Inflammatory dysregulation of blood monocytes in Parkinson’s disease patients. Acta Neuropathol. 2014;128(5):651–63.
Fujimura N, Xu B, Dalman J, et al. CCR2 inhibition sequesters multiple subsets of leukocytes in the bone marrow. Sci Rep. 2015;5:11664.
Lucot KL, Stevens MY, Bonham TA, et al. Tracking innate immune activation in a mouse model of Parkinson’s Disease using TREM1 and TSPO PET tracers. J Nucl Med. 2022;63(10):1570–8.
Birney E. Mendelian randomization. Cold Spring Harb Perspect Med. 2022;12(4): a041302.
Davey Smith G, Hemani G. Mendelian randomization: genetic anchors for causal inference in epidemiological studies. Hum Mol Genet. 2014;23(R1):R89–98.
Bowden J, Holmes MV. Meta-analysis and Mendelian randomization: a review. Res Synth Methods. 2019;10(4):486–96.
Wang C, Zhu D, Zhang D, et al. Causal role of immune cells in schizophrenia: Mendelian randomization (MR) study. BMC Psychiatry. 2023;23(1):590.
Gong Z, Liu Y, Ding F, et al. Natural killer cells-related immune traits and amyotrophic lateral sclerosis: A Mendelian randomization study. Front Neurosci. 2022;16: 981371.
Nalls MA, Blauwendraat C, Vallerga CL, et al. Identification of novel risk loci, causal insights, and heritable risk for Parkinson’s disease: a meta-analysis of genome-wide association studies. Lancet Neurol. 2019;18(12):1091–102.
Orrù V, Steri M, Sidore C, et al. Complex genetic signatures in immune cells underlie autoimmunity and inform therapy. Nat Genet. 2020;52(10):1036–45.
Sidore C, Busonero F, Maschio A, et al. Genome sequencing elucidates Sardinian genetic architecture and augments association analyses for lipid and blood inflammatory markers. Nat Genet. 2015;47(11):1272–81.
Yu XH, Yang YQ, Cao RR, et al. The causal role of gut microbiota in development of osteoarthritis. Osteoarthr Cartil. 2021;29(12):1741–50.
Auton A, Brooks LD, Durbin RM, et al. A global reference for human genetic variation. Nature. 2015;526(7571):68–74.
Burgess S, Small DS, Thompson SG. A review of instrumental variable estimators for Mendelian randomization. Stat Methods Med Res. 2017;26(5):2333–55.
Yavorska OO, Burgess S. MendelianRandomization: an R package for performing Mendelian randomization analyses using summarized data. Int J Epidemiol. 2017;46(6):1734–9.
Burgess S, Thompson SG. Interpreting findings from Mendelian randomization using the MR-Egger method. Eur J Epidemiol. 2017;32(5):377–89.
Bowden J, Davey Smith G, Haycock PC, et al. Consistent estimation in Mendelian randomization with some invalid instruments using a weighted median estimator. Genet Epidemiol. 2016;40(4):304–14.
Hartwig FP, Davey Smith G, Bowden J. Robust inference in summary data Mendelian randomization via the zero modal pleiotropy assumption. Int J Epidemiol. 2017;46(6):1985–98.
**ang M, Wang Y, Gao Z, et al. Exploring causal correlations between inflammatory cytokines and systemic lupus erythematosus: A Mendelian randomization. Front Immunol. 2022;13: 985729.
Verbanck M, Chen CY, Neale B, et al. Detection of widespread horizontal pleiotropy in causal relationships inferred from Mendelian randomization between complex traits and diseases. Nat Genet. 2018;50(5):693–8.
Szklarczyk D, Gable AL, Lyon D, et al. STRING v11: protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 2019;47(D1):D607–13.
Weiskopf K, Schnorr PJ, Pang WW, et al. Myeloid cell origins, differentiation, and clinical implications. Microbiol Spectr. 2016. https://doi.org/10.1128/microbiolspec.MCHD-0031-2016.
Thome AD, Atassi F, Wang J, et al. Ex vivo expansion of dysfunctional regulatory T lymphocytes restores suppressive function in Parkinson’s disease. NPJ Parkinsons Dis. 2021;7(1):41.
Wang B, Ma Y, Li S, et al. GSDMD in peripheral myeloid cells regulates microglial immune training and neuroinflammation in Parkinson’s disease. Acta Pharm Sin B. 2023;13(6):2663–79.
Zhang QS, Heng Y, Yuan YH, et al. Pathological α-synuclein exacerbates the progression of Parkinson’s disease through microglial activation. Toxicol Lett. 2017;265:30–7.
Cao S, Standaert DG, Harms AS. The gamma chain subunit of Fc receptors is required for alpha-synuclein-induced pro-inflammatory signaling in microglia. J Neuroinflammation. 2012;9:259.
Landoni VI, Martire-Greco D, Rodriguez-Rodrigues N, et al. Immature myeloid Gr-1+ CD11b+ cells from lipopolysaccharide-immunosuppressed mice acquire inhibitory activity in the bone marrow and migrate to lymph nodes to exert their suppressive function. Clin Sci (Lond). 2016;130(4):259–71.
Zhang Y, **e J, Han G, et al. Detection and clinical significance of myeloid-derived suppressor cells in peripheral blood of patients with rectal carcinoma. Zhonghua Wei Chang Wai Ke Za Zhi. 2017;20(7):798–802.
Kauppinen A, Kaarniranta K, Salminen A. Potential role of myeloid-derived suppressor cells (MDSCs) in age-related macular degeneration (AMD). Front Immunol. 2020;11:384.
Hu S, Li S, Ning W, et al. Identifying crosstalk genetic biomarkers linking a neurodegenerative disease, Parkinson’s disease, and periodontitis using integrated bioinformatics analyses. Front Aging Neurosci. 2022;14:1032401.
Ostanin DV, Bhattacharya D. Myeloid-derived suppressor cells in the inflammatory bowel diseases. Inflamm Bowel Dis. 2013;19(11):2468–77.
Hoechst B, Gamrekelashvili J, Manns MP, et al. Plasticity of human Th17 cells and iTregs is orchestrated by different subsets of myeloid cells. Blood. 2011;117(24):6532–41.
Yang L, Guo C, Zhu J, et al. Increased levels of pro-inflammatory and anti-inflammatory cellular responses in Parkinson’s disease patients: search for a disease indicator. Med Sci Monit. 2017;23:2972–8.
Chen CK, Wu YT, Chang YC. Periodontal inflammatory disease is associated with the risk of Parkinson’s disease: a population-based retrospective matched-cohort study. PeerJ. 2017;5: e3647.
Park J, Lee JW, Cooper SC, et al. Parkinson disease-associated LRRK2 G2019S transgene disrupts marrow myelopoiesis and peripheral Th17 response. J Leukoc Biol. 2017;102(4):1093–102.
Kim KS, Marcogliese PC, Yang J, et al. Regulation of myeloid cell phagocytosis by LRRK2 via WAVE2 complex stabilization is altered in Parkinson’s disease. Proc Natl Acad Sci U S A. 2018;115(22):E5164–73.
Li A, Peng Y, Taiclet LM, et al. Analysis of hidradenitis suppurativa-linked mutations in four genes and the effects of PSEN1-P242LfsX11 on cytokine and chemokine expression in macrophages. Hum Mol Genet. 2019;28(7):1173–82.
Tian Q, Sun X, Li C, et al. CD33 polymorphisms and Parkinson’s disease Parkinson’s disease in northern Chinese Han population: A case-control study. Neurosci Lett. 2023;812: 137400.
Siokas V, Arseniou S, Aloizou AM, et al. CD33 rs3865444 as a risk factor for Parkinson’s disease. Neurosci Lett. 2021;748: 135709.
Hakami MA, Alotaibi BS, Hazazi A, et al. Identification of potential inhibitors of tropomyosin receptor kinase B targeting CNS-related disorders and cancers. J Biomol Struct Dyn. 2023;42(6):2965–75.
Zhu G, Li J, He L, et al. MPTP-induced changes in hippocampal synaptic plasticity and memory are prevented by memantine through the BDNF-TrkB pathway. Br J Pharmacol. 2015;172(9):2354–68.
Luo D, Shi Y, Wang J, et al. 7,8-dihydroxyflavone protects 6-OHDA and MPTP induced dopaminergic neurons degeneration through activation of TrkB in rodents. Neurosci Lett. 2016;620:43–9.
Payton JE, Perrin RJ, Woods WS, et al. Structural determinants of PLD2 inhibition by alpha-synuclein. J Mol Biol. 2004;337(4):1001–9.
Mendez-Gomez HR, Singh J, Meyers C, et al. The lipase activity of phospholipase D2 is responsible for nigral neurodegeneration in a rat model of Parkinson’s Disease. Neuroscience. 2018;377:174–83.
Pan B, Niu B, He Y, et al. Integrative multilevel exploration of the mechanism by which Er-Zhi-Wan alleviates the Parkinson’s disease (PD)-like phenotype in the MPTP-induced PD mouse model. Biomed Pharmacother. 2023;165: 115021.
Alieva AK, Rudenok MM, Novosadova EV, et al. Whole-transcriptome analysis of dermal fibroblasts, derived from three pairs of monozygotic twins, discordant for Parkinson’s Disease. J Mol Neurosci. 2020;70(2):284–93.
Mishra A, Malik R, Hachiya T, et al. Stroke genetics informs drug discovery and risk prediction across ancestries. Nature. 2022;611(7934):115–23.
Abdelmoaty MM, Machhi J, Yeapuri P, et al. Monocyte biomarkers define sargramostim treatment outcomes for Parkinson’s disease. Clin Transl Med. 2022;12(7): e958.
Olson KE, Namminga KL, Lu Y, et al. Safety, tolerability, and immune-biomarker profiling for year-long sargramostim treatment of Parkinson’s disease. EBioMedicine. 2021;67: 103380.
Acknowledgements
This study was supported by Jilin Province Center for Precision Medicine Diagnosis and Treatment of Nervous System Diseases (No. 20200602045ZP), and Development of human mesenchymal stem cell exosome-loaded astaxanthin preparations and its preclinical study for the treatment of Parkinson's disease. This research was also funded by Norman Bethune Program of Jilin University (No. 2022B28) and Health Research Talent Program of Jilin Province (No. 2022SC234).
Author information
Authors and Affiliations
Contributions
Jiajun Chen designed the study and was considered corresponding author. Wei Quan mainly completes article writing and data analysis, and was listed as the first author. Yidan Qin participated in analysis data. The remaining authors were involved in article guidance, revision, etc. All authors read and approve the final paper.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Responsible Editor: John Di Battista.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Quan, W., Qin, Y., Li, J. et al. Causal role of myeloid cells in Parkinson’s disease: Mendelian randomization study. Inflamm. Res. 73, 809–818 (2024). https://doi.org/10.1007/s00011-024-01867-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00011-024-01867-8