Abstract
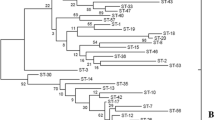
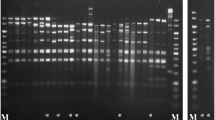
Diversity in the ruminal bacterial speciesSelenomonas ruminantium has been investigated by DNA fingerprinting, DNA-DNA hybridization, plasmid analysis, bacteriophage sensitivity, and monoclonal antibody-based immunoassay. Twenty different isolates from the sheep rumen were initially classified morphologically and by carbon source utilization. DNA fingerprint analyses and quantitative genomic DNA hybridizations showed that limited grou** of these isolates was possible, with the largest group comprising four isolates, and two other groups comprising two isolates each. The remaining isolates were unique. Plasmids in four different size classes, 2.5, 3.7, 6.5 and 12.0 kbp, were identified, but these did not appear in all isolates. There was no apparent relationship between DNA fingerprint pattern and plasmid content. Only three isolates were sensitive to theS. ruminantium-specific temperate bacteriophage S-1. These data indicate that substantial genetic diversity exists within the ruminal speciesS. ruminantium, but that at least one strain may represent up to 20% of isolates.
Similar content being viewed by others
Literature Cited
Brooker JD, Stokes B (1990) Monoclonal antibodies against the rumen bacterial species.Selenomonas ruminantium. Appl Environ Microbiol 56:2193–2199
Bryant MP (1977) Microbiology of the rumen. In: Swanson MJ (ed) Dukes physiology of domestic animals. Ithaca, NY: Cornell University Press, pp 287–304
Bryant MP, Burkey LA (1953) Culture method and some characteristics of some of the more numerous groups of bacteria in the bovine rumen J Dairy Sci 36:305–217
Cooper DN, Schmidtke J (1984) DNA restriction fragment length polymorphism and heterozygosity in the human genome. Human Genet 66:1–16
Dean RG, Martin SA, Carver C (1989) Isolation of plasmid DNA from the ruminal bacteriumSelenomonas ruminantium HD4. Lett Appl Microbiol 8:45–48
Feinberg AP, Vogelstein B (1984) A technique for radiolabelling DNA restriction endonuclease fragments to high specific activity. Anal Biochem 137:226–267
Flint HJ, Duncan SH, Bisset J, Steward CS (1988) The isolation of tetracycline resistant strains of strictly anaerobic bacteria from the rumen. Lett Appl Microbiol 6:113–115
Grimont PAD (1988) Use of DNA reassociation in bacterial classification. Can J Microbiol 34:541–545
Hazlewood GP, Theodorou MK, Hutchings A, Jordan JL, Galfre G (1986) Preparation and characterization of monoclonal antibodies to aButrivibrio sp. and their potential use in the identification of rumenButyrivibrios using and ELISA. J Gen Microbiol 132:43–52
Hudman JF, Gregg K (1989) Genetic diversity among strains of bacteria from the rumen. Curr Microbiol 19:313–318
Lockington RA, Attwood GT, Brooker JD (1988) The isolation and characterization of a temperate bacteriophage from the rumen anaerobe,Selenomonas ruminantium. Appl Environ Microbiol 54:1575–1580
Macario AJL, Conway de Macario E (1988) Quantitative immunologic analysis of the methanogenic flora of digestors reveals a considerable diversity. Appl Environ Microbiol 54:79–86
Ogimoto K, Imai S (1984) Atlas of rumen microbiology. Tokyo: Japan Scientific Societies Press
Tiwari AD, Bryant MP, Wolfe RS (1968) Simple method for isolation ofS. ruminantium and some nutritional characteristics of the species. J Dairy Sci 52:2054–2056
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Ning, Z., Attwood, G.T., Lockington, R.A. et al. Genetic diversity in ruminal isolates ofSelenomonas ruminantium . Current Microbiology 22, 279–284 (1991). https://doi.org/10.1007/BF02091955
Issue Date:
DOI: https://doi.org/10.1007/BF02091955